Fitting CaMKII parameters to experimental decay rates.
using Model
using Model: second, Hz
using ADTypes
using CurveFit
using DiffEqCallbacks
using DifferentialEquations
using ForwardDiff
using LinearAlgebra
using ModelingToolkit
using ModelingToolkit: t_nounits as t, D_nounits as D
using Optimization
using OptimizationOptimJL
using OrdinaryDiffEqSDIRK
using PlotsExperimental data¶
CaMKII activity decay time scales after 1Hz pacing for 15.0, 30.0, 60.0, 90.0 seconds.
pacing_durations = [15.0, 30.0, 60.0, 90.0]
experimental_taus = [16.48, 16.73, 17.65, 18.08]4-element Vector{Float64}:
16.48
16.73
17.65
18.08Setup problem¶
stimstart = 30.0second
stimend = 130.0second
tend = 205.0second
alg = KenCarp4()
@time "Building ODE system" sys = Model.DEFAULT_SYS
@time "Building ODE problem" prob = ODEProblem(sys, [sys.r_CaMK=>10/second], tend)Building ODE system: 0.000023 seconds (48 allocations: 2.203 KiB)
Building ODE problem: 63.102725 seconds (115.81 M allocations: 5.690 GiB, 4.68% gc time, 99.79% compilation time: 75% of which was recompilation)
ODEProblem with uType Vector{Float64} and tType Float64. In-place: true
Initialization status: FULLY_DETERMINED
Non-trivial mass matrix: false
timespan: (0.0, 205000.0)
u0: 75-element Vector{Float64}:
830.0
830.0
0.0026
0.07192
0.07831
0.26081
0.00977
0.00188
0.09243
0.22156
⋮
0.12113
0.12113
0.12113
0.12113
0.12113
0.12113
150952.75035000002
13838.37602
-68.79268@unpack Istim, CaMKAct = sys
pace15 = Model.build_stim_callbacks(Istim, stimstart + 15 * second; period=1second, starttime=stimstart)
pace30 = Model.build_stim_callbacks(Istim, stimstart + 30 * second; period=1second, starttime=stimstart)
pace60 = Model.build_stim_callbacks(Istim, stimstart + 60 * second; period=1second, starttime=stimstart)
pace90 = Model.build_stim_callbacks(Istim, stimstart + 90 * second; period=1second, starttime=stimstart);Test run with default parameters¶
@time "Solve problems" sols = map([pace15, pace30, pace60, pace90]) do cb
solve(prob, alg, callback = cb)
end
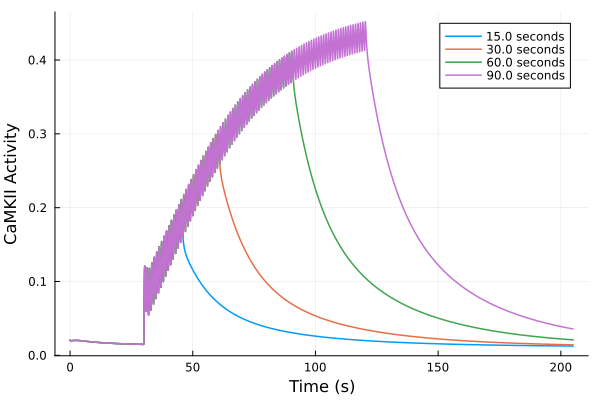
plot()
for (sol, dur) in zip(sols, pacing_durations)
plot!(sol, idxs=(t/1000, CaMKAct), label="$(dur) seconds")
end
plot!(xlabel="Time (s)", ylabel="CaMKII Activity", title="", legend=:topright)Solve problems: 2.375672 seconds (1.37 M allocations: 91.876 MiB, 31.73% compilation time: 85% of which was recompilation)
Loss function¶
Changing CaMKII parameters to fit decay rate in the experiments.
@unpack kphos_CaMK, kdeph_CaMK, kb_CaMKP, k_P1_P2, k_P2_P1, CaMKAct, r_CaMK = sys
kphos_dephos_ratio = prob.ps[kphos_CaMK] / prob.ps[kdeph_CaMK]
p1p2_ratio = prob.ps[k_P2_P1] / prob.ps[k_P1_P2]
data = (
prob=prob,
cbs=[pace15, pace30, pace60, pace90],
experimental_taus=experimental_taus,
pacing_durations=pacing_durations,
kphos_dephos_ratio=kphos_dephos_ratio,
p1p2_ratio = p1p2_ratio,
stimstart = stimstart,
tend=tend
);
function loss(theta, data)
@unpack prob, cbs, experimental_taus, pacing_durations, kphos_dephos_ratio, p1p2_ratio, stimstart, tend = data
@unpack kdeph_CaMK, kphos_CaMK, kb_CaMKP, k_P1_P2, k_P2_P1, CaMKAct, r_CaMK = prob.f.sys
dephos_rate = exp10(theta[1])
kb2_rate = exp10(theta[2])
k_P1_P2_rate = exp10(theta[3])
k_P2_P1_rate = k_P1_P2_rate * p1p2_ratio
r_CaMK_rate = exp10(theta[4])
# Parallel ensemble simulation
function prob_func(prob, ctx)
i = ctx.sim_id
remake(prob, p=[
kdeph_CaMK => dephos_rate,
kphos_CaMK => kphos_dephos_ratio * dephos_rate,
kb_CaMKP => kb2_rate,
k_P1_P2 => k_P1_P2_rate,
k_P2_P1 => k_P2_P1_rate,
r_CaMK => r_CaMK_rate
],
callback=cbs[i],
saveat=(stimstart + pacing_durations[i] * second):1second:tend
)
end
# Calculate loss in the output function
function output_func(sol, ctx)
SciMLBase.successful_retcode(sol) || return (Inf, false)
i = ctx.sim_id
stimend = stimstart + pacing_durations[i] * second
ts = collect(range(0.0, stop=50.0, step=5.0))
ysim = sol(stimend .+ ts .* second, idxs=CaMKAct).u
fit = solve(CurveFitProblem(ts, ysim), ExpSumFitAlgorithm(n=1, withconst=true))
tau = inv(-fit.u.λ[])
tauexpected = experimental_taus[i]
l2 = (tau - tauexpected)^2
return (l2, false)
end
ensemble_prob = EnsembleProblem(prob; prob_func, output_func)
sim = solve(ensemble_prob, alg, EnsembleThreads(); trajectories=length(pacing_durations), maxiters=10000)
return sum(sim)
endloss (generic function with 1 method)Test the loss function
theta = [log10(prob.ps[kdeph_CaMK]), log10(prob.ps[kb_CaMKP]), log10(prob.ps[k_P1_P2]), log10(prob.ps[r_CaMK])]
@time loss(theta, data) 12.785629 seconds (35.24 M allocations: 1.799 GiB, 7.47% gc time, 27553 lock conflicts, 144.18% compilation time: <1% of which was recompilation)
1.3401087281337494Optimization¶
optf = OptimizationFunction(loss)
optprob = OptimizationProblem(optf, theta, data, lb=[-1, -1, -1, 0] + theta, ub=[1, 1, 1, 1] + theta)
optalg = Optim.SAMIN()
maxtime = haskey(ENV, "JULIA_CI") ? 60 : 2000
@time fitted_dephos = solve(optprob, optalg; maxiters=5000, maxtime=maxtime)
@show fitted_dephos.objective
fitted_dephos.stats 61.767105 seconds (177.88 M allocations: 11.723 GiB, 13.48% gc time, 562270 lock conflicts, 2.69% compilation time)
fitted_dephos.objective = 0.09972371028067181
SciMLBase.OptimizationStats
Number of iterations: 30
Time in seconds: 61.024437
Number of function evaluations: 30
Number of gradient evaluations: 0
Number of hessian evaluations: 0params = Dict(kdeph_CaMK => exp10(fitted_dephos.u[1]), kphos_CaMK => data.kphos_dephos_ratio * exp10(fitted_dephos.u[1]), kb_CaMKP => exp10(fitted_dephos.u[2]), k_P1_P2 => exp10(fitted_dephos.u[3]), k_P2_P1 => exp10(fitted_dephos.u[3]) * p1p2_ratio, r_CaMK => exp10(fitted_dephos.u[4]))
println("Fitted parameters:")
println("Dephosphorylation time: " , 1e-3 / params[kdeph_CaMK], " seconds.")
println("Dephosphorylation rate: " , params[kdeph_CaMK] * 1000, " Hz.")
println("Autophosphorylation rate of CaMKB: " , params[kphos_CaMK] * 1000, " Hz")
println("CaM dissociation time of CaMKP: " , 1e-3 / params[kb_CaMKP], " seconds.")
println("CaM dissociation rate of CaMKP: " , params[kb_CaMKP] * 1000, " Hz.")
println("P1 to P2 time: " , 1e-3 / params[k_P1_P2], " seconds.")
println("P1 to P2 rate: " , params[k_P1_P2] * 1000, " Hz.")
println("P2 to P1 time: " , 1e-3 / params[k_P2_P1], " seconds.")
println("P2 to P1 rate: " , params[k_P2_P1] * 1000, " Hz.")
println("CaMK binding rate: " , params[r_CaMK] * 1000, " Hz.")Fitted parameters:
Dephosphorylation time: 12.013799999999998 seconds.
Dephosphorylation rate: 0.08323761008173935 Hz.
Autophosphorylation rate of CaMKB: 1.8438000000000003 Hz
CaM dissociation time of CaMKP: 0.09251378254195453 seconds.
CaM dissociation rate of CaMKP: 10.809200235073138 Hz.
P1 to P2 time: 114.61420886542861 seconds.
P1 to P2 rate: 0.00872492171694109 Hz.
P2 to P1 time: 28.657744917112005 seconds.
P2 to P1 rate: 0.03489458095507312 Hz.
CaMK binding rate: 73.67208218821155 Hz.
Test fitted parameters¶
newprob = remake(prob, p=[kdeph_CaMK => params[kdeph_CaMK], kphos_CaMK => params[kphos_CaMK], kb_CaMKP => params[kb_CaMKP], k_P1_P2 => params[k_P1_P2], k_P2_P1 => params[k_P2_P1], r_CaMK => params[r_CaMK]])
sols = map([pace15, pace30, pace60, pace90]) do cb
solve(newprob, alg, callback = cb)
end;
plot()
for (sol, dur) in zip(sols, [15.0, 30.0, 60.0, 90.0])
plot!(sol, idxs=(t/1000, CaMKAct), label="$(dur) seconds")
end
plot!(xlabel="Time (s)", ylabel="Active CaMKII fraction", title="Pacing durations", legend=:topright)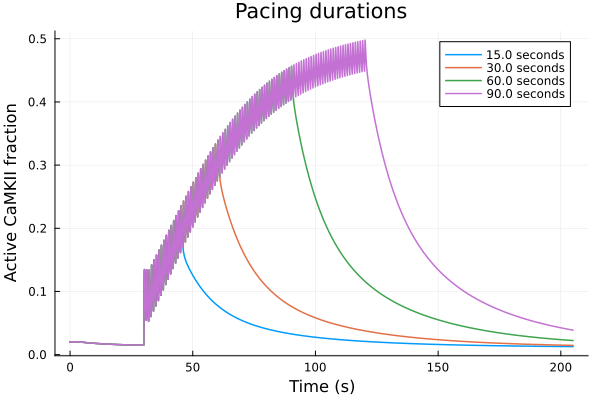
Decay rates¶
Fit data from simulations against an exponential decay model. Record 50 seconds after pacing ends.
ts = collect(range(0.0, stop=50.0, step=5.0)) ## in seconds
stimstart = 30.0second
ysim_15 = sols[1](stimstart+15second:5second:stimstart+15second+50second ; idxs=sys.CaMKAct).u
ysim_30 = sols[2](stimstart+30second:5second:stimstart+30second+50second ; idxs=sys.CaMKAct).u
ysim_60 = sols[3](stimstart+60second:5second:stimstart+60second+50second ; idxs=sys.CaMKAct).u
ysim_90 = sols[4](stimstart+90second:5second:stimstart+90second+50second ; idxs=sys.CaMKAct).u
fit_sim_15 = solve(CurveFitProblem(ts, ysim_15), ExpSumFitAlgorithm(n=1, withconst=true))
fit_sim_30 = solve(CurveFitProblem(ts, ysim_30), ExpSumFitAlgorithm(n=1, withconst=true))
fit_sim_60 = solve(CurveFitProblem(ts, ysim_60), ExpSumFitAlgorithm(n=1, withconst=true))
fit_sim_90 = solve(CurveFitProblem(ts, ysim_90), ExpSumFitAlgorithm(n=1, withconst=true))retcode: Success
alg: ExpSumFitAlgorithm
residuals mean: 2.6493958542191237e-17
u: [0.057516292769088415, 0.3908368331517097, -0.05521880920512651]Fitting results (simulations)¶
p1s = plot(ts, ysim_15, label="Sim 15 sec")
plot!(p1s, ts, predict(fit_sim_15), label="Fit", linestyle=:dash)
p2s = plot(ts, ysim_30, label="Sim 30 sec")
plot!(p2s, ts, predict(fit_sim_30), label="Fit", linestyle=:dash)
p3s = plot(ts, ysim_60, label="Sim 60 sec")
plot!(p3s, ts, predict(fit_sim_60), label="Fit", linestyle=:dash)
p4s = plot(ts, ysim_90, label="Sim 90 sec")
plot!(p4s, ts, predict(fit_sim_90), label="Fit", linestyle=:dash)
plot(p1s, p2s, p3s, p4s, layout=(2,2), xlabel="Time (s)", ylabel="Active CaMKII fraction")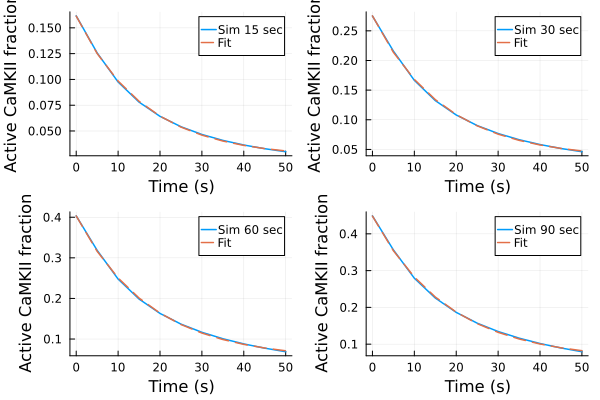
Decay time scales (tau)¶
tau_sim_15 = inv(-fit_sim_15.u.λ[])
tau_sim_30 = inv(-fit_sim_30.u.λ[])
tau_sim_60 = inv(-fit_sim_60.u.λ[])
tau_sim_90 = inv(-fit_sim_90.u.λ[])
println("The time scales for simulations: ")
for (tau, dur) in zip((tau_sim_15, tau_sim_30, tau_sim_60, tau_sim_90), (15, 30, 60, 90))
println("$dur sec pacing is $(round(tau; digits=2)) seconds.")
endThe time scales for simulations:
15 sec pacing is 16.26 seconds.
30 sec pacing is 16.95 seconds.
60 sec pacing is 17.62 seconds.
90 sec pacing is 18.11 seconds.
This notebook was generated using Literate.jl.