Caffeine increase RyR opening sensitivity to luminal and subspace calcium. In this model, we decrease the mid saturation sub-SR calcium concentration for the opening rate.
using Model
using Model: second
using CSV
using DataFrames
using DiffEqCallbacks
using DifferentialEquations
using ModelingToolkit
using OrdinaryDiffEqSDIRK
using Plots
using SteadyStateDiffEq
import Dates
Plots.default(lw=1.5)@time "Build system" sys = Model.DEFAULT_SYS
tend = 500second
@time "Build problem" prob = ODEProblem(sys, [], tend)
stimstart = 100second
stimend = 300second
alg = KenCarp4()
function add_coffee_affect!(integrator)
integrator.ps[sys.RyRsensitivity] = 10
endBuild system: 0.000025 seconds (38 allocations: 1.766 KiB)
Build problem: 35.937021 seconds (56.50 M allocations: 2.769 GiB, 5.19% gc time, 99.40% compilation time: 10% of which was recompilation)
add_coffee_affect! (generic function with 1 method)@unpack Istim = sys
callback = build_stim_callbacks(Istim, stimend; period=1second, starttime=stimstart)SciMLBase.CallbackSet{Tuple{}, Tuple{SciMLBase.DiscreteCallback{DiffEqCallbacks.PresetTimeFunction{StepRange{Int64, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#20#21"{Float64, Float64, Symbolics.Num}}, Model.var"#20#21"{Float64, Float64, Symbolics.Num}, DiffEqCallbacks.PresetTimeFunction{StepRange{Int64, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#20#21"{Float64, Float64, Symbolics.Num}}, typeof(SciMLBase.FINALIZE_DEFAULT), Nothing, Tuple{}}, SciMLBase.DiscreteCallback{DiffEqCallbacks.PresetTimeFunction{StepRangeLen{Float64, Base.TwicePrecision{Float64}, Base.TwicePrecision{Float64}, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#22#23"{Float64, Float64, Symbolics.Num}}, Model.var"#22#23"{Float64, Float64, Symbolics.Num}, DiffEqCallbacks.PresetTimeFunction{StepRangeLen{Float64, Base.TwicePrecision{Float64}, Base.TwicePrecision{Float64}, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#22#23"{Float64, Float64, Symbolics.Num}}, typeof(SciMLBase.FINALIZE_DEFAULT), Nothing, Tuple{}}}}((), (SciMLBase.DiscreteCallback{DiffEqCallbacks.PresetTimeFunction{StepRange{Int64, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#20#21"{Float64, Float64, Symbolics.Num}}, Model.var"#20#21"{Float64, Float64, Symbolics.Num}, DiffEqCallbacks.PresetTimeFunction{StepRange{Int64, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#20#21"{Float64, Float64, Symbolics.Num}}, typeof(SciMLBase.FINALIZE_DEFAULT), Nothing, Tuple{}}(DiffEqCallbacks.PresetTimeFunction{StepRange{Int64, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#20#21"{Float64, Float64, Symbolics.Num}}(100000:1000:300000, true, SciMLBase.INITIALIZE_DEFAULT, Model.var"#20#21"{Float64, Float64, Symbolics.Num}(-80.0, 0.5, Istim)), Model.var"#20#21"{Float64, Float64, Symbolics.Num}(-80.0, 0.5, Istim), DiffEqCallbacks.PresetTimeFunction{StepRange{Int64, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#20#21"{Float64, Float64, Symbolics.Num}}(100000:1000:300000, true, SciMLBase.INITIALIZE_DEFAULT, Model.var"#20#21"{Float64, Float64, Symbolics.Num}(-80.0, 0.5, Istim)), SciMLBase.FINALIZE_DEFAULT, Bool[1, 1], nothing, (), true), SciMLBase.DiscreteCallback{DiffEqCallbacks.PresetTimeFunction{StepRangeLen{Float64, Base.TwicePrecision{Float64}, Base.TwicePrecision{Float64}, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#22#23"{Float64, Float64, Symbolics.Num}}, Model.var"#22#23"{Float64, Float64, Symbolics.Num}, DiffEqCallbacks.PresetTimeFunction{StepRangeLen{Float64, Base.TwicePrecision{Float64}, Base.TwicePrecision{Float64}, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#22#23"{Float64, Float64, Symbolics.Num}}, typeof(SciMLBase.FINALIZE_DEFAULT), Nothing, Tuple{}}(DiffEqCallbacks.PresetTimeFunction{StepRangeLen{Float64, Base.TwicePrecision{Float64}, Base.TwicePrecision{Float64}, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#22#23"{Float64, Float64, Symbolics.Num}}(100000.5:1000.0:300000.5, true, SciMLBase.INITIALIZE_DEFAULT, Model.var"#22#23"{Float64, Float64, Symbolics.Num}(0.0, 0.5, Istim)), Model.var"#22#23"{Float64, Float64, Symbolics.Num}(0.0, 0.5, Istim), DiffEqCallbacks.PresetTimeFunction{StepRangeLen{Float64, Base.TwicePrecision{Float64}, Base.TwicePrecision{Float64}, Int64}, typeof(SciMLBase.INITIALIZE_DEFAULT), Model.var"#22#23"{Float64, Float64, Symbolics.Num}}(100000.5:1000.0:300000.5, true, SciMLBase.INITIALIZE_DEFAULT, Model.var"#22#23"{Float64, Float64, Symbolics.Num}(0.0, 0.5, Istim)), SciMLBase.FINALIZE_DEFAULT, Bool[1, 1], nothing, (), true)))Add caffeine at t = 200 econd
callback_caf = CallbackSet(build_stim_callbacks(Istim, stimend; period=1second, starttime=stimstart), PresetTimeCallback(200.0second, add_coffee_affect!));Single-dose caffeine¶
@time sol = solve(prob, alg; callback)
@time sol_caf = solve(prob, alg; callback=callback_caf)
i = (sys.t / 1000, sys.vm)
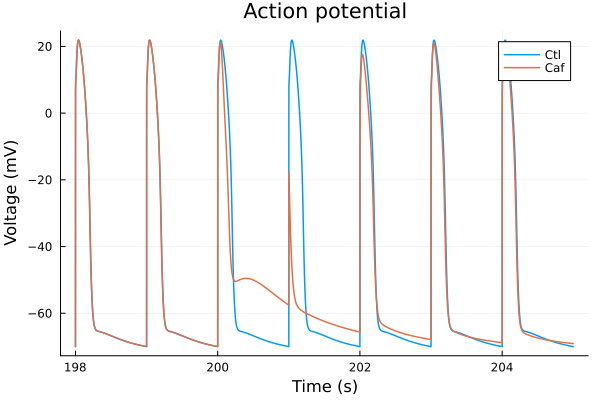
plot(sol, idxs=i, title="Action potential", lab="Ctl", tspan=(198second, 205second))
plot!(sol_caf, idxs=i, lab="Caf", tspan=(198second, 205second), ylabel="Voltage (mV)", xlabel="Time (s)") 8.795650 seconds (15.52 M allocations: 723.184 MiB, 8.37% gc time, 75.21% compilation time)
4.271983 seconds (3.46 M allocations: 192.990 MiB, 1.32% gc time, 51.84% compilation time)
i = (sys.t / 1000, sys.Cai_sub_SR * 1000)
plot(sol, idxs=i, title="Calcium transient (During caffeine addition)", lab="Ctl", tspan=(198second, 205second))
plot!(sol_caf, idxs=i, tspan=(198second, 205second), lab="Caf", ylabel="Subspace calcium (nM)", xlabel="Time (s)")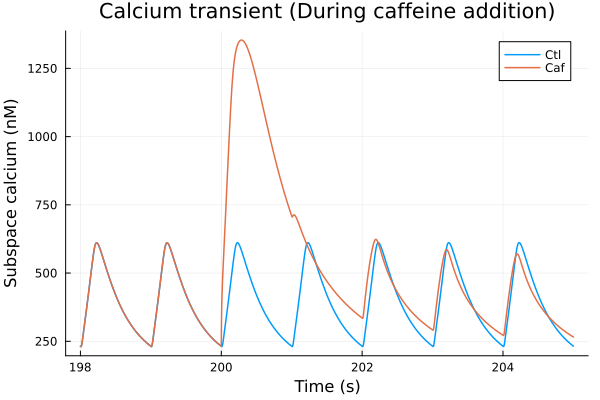
i = (sys.t / 1000, sys.PO1RyR)
plot(sol, idxs=i, title="RyR open (During caffeine addition)", lab="Ctl", tspan=(198second, 205second))
plot!(sol_caf, idxs=i, tspan=(198second, 205second), lab="Caf", ylabel="Open probability", ylims=(0, 1), xlabel="Time (s)")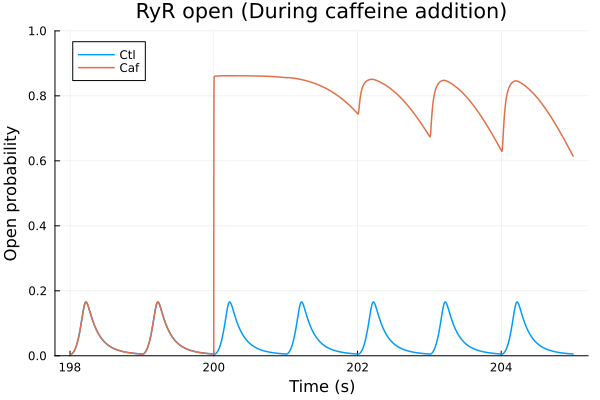
i = (sys.t / 1000, sys.Cai_sub_SR * 1000)
plot(sol, idxs=i, title="Calcium transient (After caffeine addition)", lab="Ctl", ylabel="Subspace calcium (nM)", tspan=(198second, 205second))
plot!(sol_caf, idxs=i, lab="Caf", xlabel="Time (s)", tspan=(198second, 205second))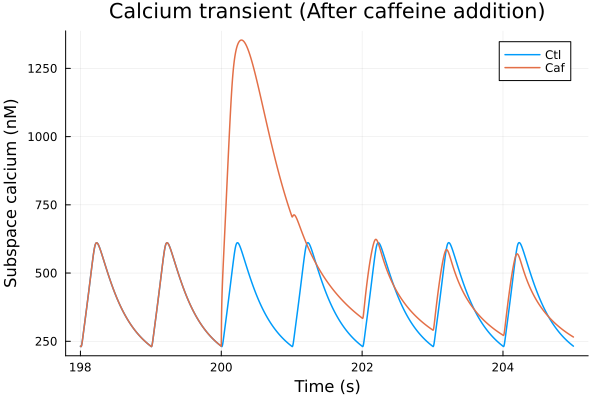
i = (sys.t / 1000, sys.CaJSR)
plot(sol, idxs=i, title="SR Calcium (During caffeine addition)", lab="Ctl", ylabel="SR calcium (μM)", tspan=(198second, 205second))
plot!(sol_caf, idxs=i, tspan=(198second, 205second), lab="Caf", ylims=(0, 850), xlabel="Time (s)")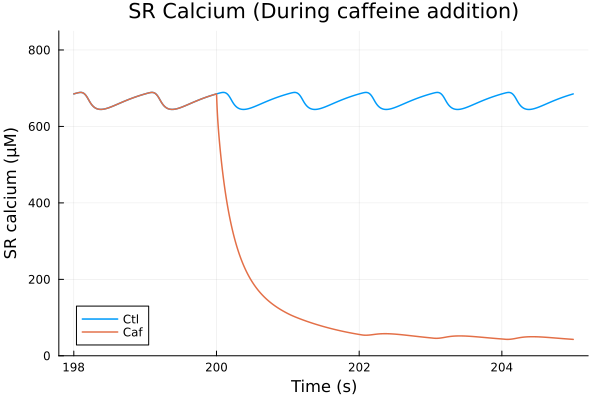
i = (sys.t / 1000, sys.Jrel)
plot(sol, idxs=sys.Jrel, title="Ca flux", lab="Ctl (Jrel)", tspan=(198second, 205second))
plot!(sol_caf, idxs=sys.Jrel, lab="Caf (Jrel)", tspan=(198second, 205second), ylabel="μM/ms", xlabel="Time (s)")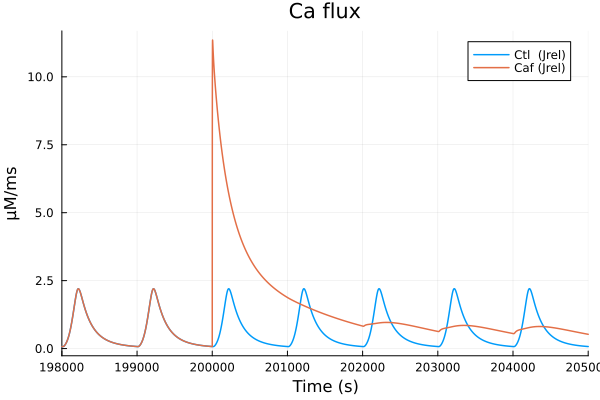
i = (sys.t / 1000, sys.CaMKAct)
plot(sol, idxs=i, title="Active CaMKII", lab="Ctl")
plot!(sol_caf, idxs=i, lab="Caf", ylabel="Active CaMKII fraction", xlabel="Time (s)")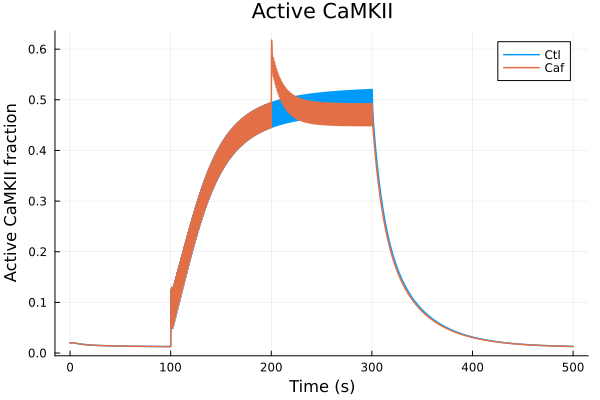
Caffeine and electrophysiology¶
Adding caffeine in the beginning of the simulation.
Adding caffeine and nifedipine in the beginning of the simulation (nifedipine blocks 90% of LCCs).
@time "Build system" sys = Model.DEFAULT_SYS
tend = 205second
stimstart = 30second
stimend = 120second
alg = KenCarp4()
@unpack Istim = sys
callback = build_stim_callbacks(Istim, stimend; period=1second, starttime=stimstart)
@time prob = ODEProblem(sys, [], tend)
@time prob_caf = ODEProblem(sys, [sys.RyRsensitivity => 10], tend)
gCaL = prob.ps[sys.GCaL]
@time prob_nif_caf = ODEProblem(sys, [sys.RyRsensitivity => 10, sys.GCaL => 0.1 * gCaL], tend)
ssalg = DynamicSS(alg)
sprob_caf = SteadyStateProblem(prob_caf)
sssol = solve(sprob_caf, ssalg; abstol=1e-10, reltol=1e-10)
@time sol = solve(prob, alg; callback)
@time sol_caf = solve(remake(prob_caf, u0=sssol.u), alg; callback)
@time sol_nif_caf = solve(remake(prob_nif_caf, u0=sssol.u), alg; callback)
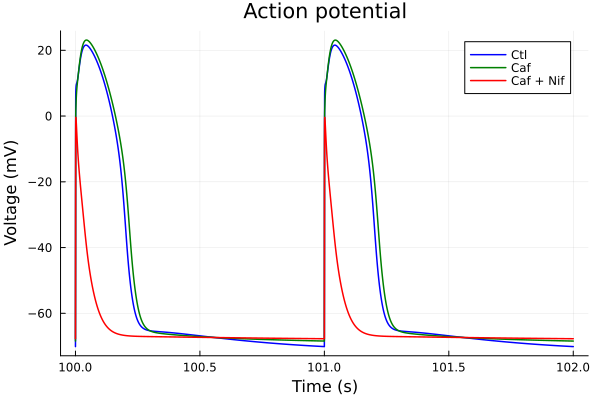
i = (sys.t / 1000, sys.vm)
tspan = (100second, 102second)
plot(sol, idxs=i, title="Action potential", lab="Ctl", color=:blue; tspan)
plot!(sol_caf, idxs=i, lab="Caf", color=:green; tspan)
plot!(sol_nif_caf, idxs=i, lab="Caf + Nif", color=:red, tspan=tspan, ylabel="Voltage (mV)", xlabel="Time (s)")Build system: 0.000010 seconds (8 allocations: 432 bytes)
0.185011 seconds (816.92 k allocations: 37.598 MiB, 28.90% compilation time)
0.285922 seconds (782.42 k allocations: 37.325 MiB, 57.05% compilation time)
0.504181 seconds (1.11 M allocations: 53.321 MiB, 74.67% compilation time)
0.983241 seconds (49.30 k allocations: 10.730 MiB, 6.57% gc time)
1.149550 seconds (380.79 k allocations: 27.363 MiB, 15.43% compilation time)
0.608752 seconds (41.54 k allocations: 8.110 MiB)
savefig("caf-ap.pdf")
savefig("caf-ap.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/caf-ap.png"i = (sys.t / 1000, sys.Cai_sub_SR * 1000)
tspan = (100second, 102second)
plot(sol, idxs=i, title="Calcium transient", lab="Ctl", color=:blue; tspan)
plot!(sol_caf, idxs=i, lab="Caf", color=:green; tspan)
plot!(sol_nif_caf, idxs=i, lab="Caf + Nif", color=:red, ylabel="Subspace calcium (nM)", xlabel="Time (s)"; tspan)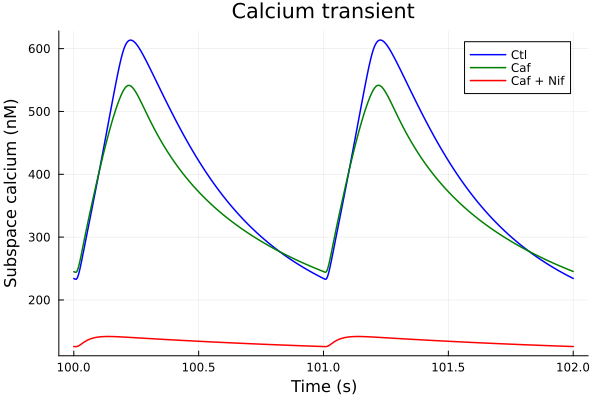
savefig("caf-cat.pdf")
savefig("caf-cat.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/caf-cat.png"i = (sys.t / 1000, sys.CaJSR)
tspan = (100second, 102second)
plot(sol, idxs=i, title="SR Calcium", lab="Ctl", ylabel="SR calcium (μM)", color=:blue; tspan)
plot!(sol_caf, idxs=i, lab="Caf", color=:green; tspan)
plot!(sol_nif_caf, idxs=i, lab="Caf + Nif", color=:red, ylims=(0, 850), xlabel="Time (s)"; tspan)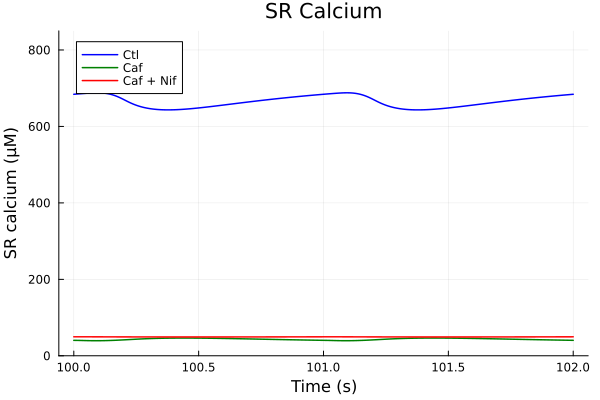
i = (sys.t / 1000, sys.CaMKAct)
plot(sol, idxs=i, title="Simulation", lab="Ctl", color=:blue)
plot!(sol_caf, idxs=i, lab="Caf", color=:green)
plot!(sol_nif_caf, idxs=i, lab="Caf + Nif", color=:red, ylabel="Active CaMKII fraction", xlabel="Time (s)")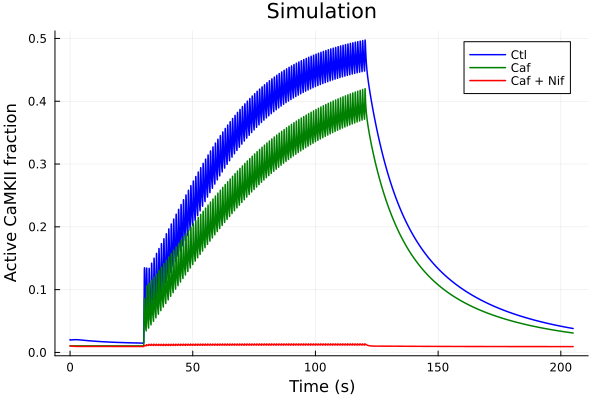
savefig("caf-camkact.pdf")
savefig("caf-camkact.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/caf-camkact.png"Experiments¶
chemicaldf = CSV.read(joinpath(@__DIR__, "data/CaMKAR-chemical.csv"), DataFrame)
ts = Dates.value.(chemicaldf[!, "Time"]) ./ 10^9
ctl = chemicaldf[!, "Ctrl Mean"]
ctl_error = chemicaldf[!, "Ctrl SD"] ./ sqrt.(chemicaldf[!, "Ctrl N"])
caf = chemicaldf[!, "caffeine 20mM Mean"]
caf_error = chemicaldf[!, "caffeine 20mM SD"] ./ sqrt.(chemicaldf[!, "caffeine 20mM N"])
plot(ts, ctl, yerr=ctl_error, lab="Control", color=:blue, markerstrokecolor=:blue)
plot!(ts, caf, yerr=caf_error, lab="Caffeine 20mM", color=:red, markerstrokecolor=:red)
plot!(xlabel="Time (s)", ylabel="CaMKII activity (A.U.)", title= "Experiment")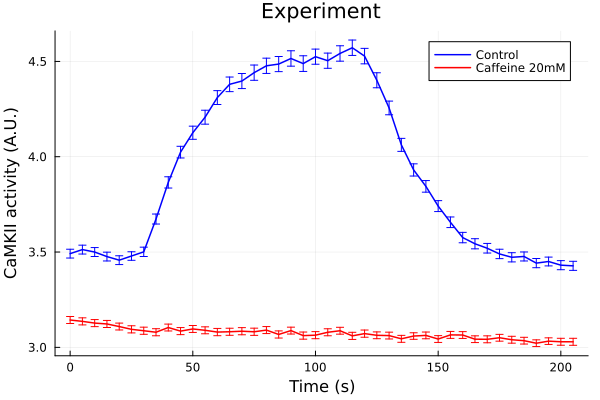
savefig("caf-exp.pdf")
savefig("caf-exp.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/caf-exp.png"This notebook was generated using Literate.jl.