Experiments vs simulations.
using Model
using Model: second, Hzusing CSV
using CurveFit
using DataFrames
using DiffEqCallbacks
using DifferentialEquations
using ModelingToolkit
using ModelingToolkit: t_nounits as t
using OrdinaryDiffEqSDIRK
using Plots
using SteadyStateDiffEq
Plots.default(lw=1.5)Experiments¶
30 sec resting + N sec 1Hz pacing + resting.
durationdf = CSV.read(joinpath(@__DIR__, "data/CaMKAR-duration.csv"), DataFrame)
ts = durationdf[!, "Time(sec)"]
fifteen = durationdf[!, "1Hz 15sec (Mean)"]
fifteen_error = durationdf[!, "1Hz 15sec (SD)"] ./ sqrt.(durationdf[!, "1Hz 15sec (N)"])
thirty = durationdf[!, "1Hz 30sec (Mean)"]
thirty_error = durationdf[!, "1Hz 30sec (SD)"] ./ sqrt.(durationdf[!, "1Hz 30sec (N)"])
sixty = durationdf[!, "1Hz 60sec (Mean)"]
sixty_error = durationdf[!, "1Hz 60sec (SD)"] ./ sqrt.(durationdf[!, "1Hz 60sec (N)"])
ninety = durationdf[!, "1Hz 90sec (Mean)"]
ninety_error = durationdf[!, "1Hz 90sec (SD)"] ./ sqrt.(durationdf[!, "1Hz 90sec (N)"])42-element Vector{Float64}:
0.07322025867393339
0.06288123803690504
0.06438696849626417
0.07094952584857149
0.0722214908336562
0.07401047293769683
0.06323601113333674
0.07088528768397788
0.06905459448562148
0.06667878213772092
⋮
0.06219021582828689
0.06524406587899219
0.05082442404251369
0.06307382411198
0.07385218047440929
0.062460484843557096
0.03877222159045237
0.05170103726195504
0.042024632430084405Note that 30 sec timeseries have a lower baseline and 90 sec timeseries have a higher baseline before pacing.
plot(ts, fifteen, yerr=fifteen_error, lab="15 sec", color=:blue, markerstrokecolor=:blue)
plot!(ts, thirty, yerr=thirty_error, lab="30 sec", color=:red, markerstrokecolor=:red)
plot!(ts, sixty, yerr=sixty_error, lab="60 sec", color=:orange, markerstrokecolor=:orange)
plot!(ts, ninety, yerr=ninety_error, lab="90 sec", color=:green, markerstrokecolor=:green)
plot!(title="Experiment", xlabel="Time (s)", ylabel="CaMKII activity (AU)")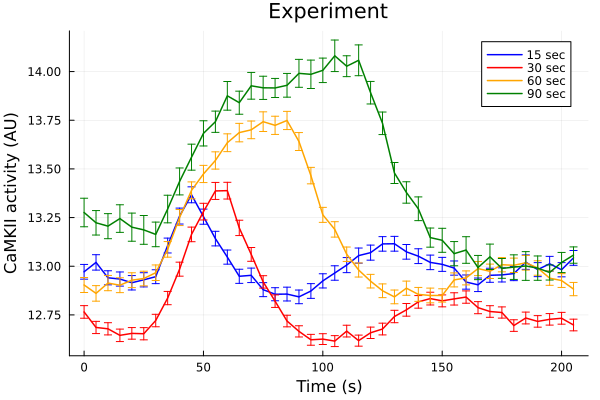
savefig("pacing-duration-exp.pdf")
savefig("pacing-duration-exp.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/pacing-duration-exp.png"Simulation¶
@time "Build system" sys = Model.DEFAULT_SYS
tend = 210second
@time "Build problem" prob = ODEProblem(sys, [], (0second, tend))
alg = KenCarp4()Build system: 0.000036 seconds (48 allocations: 2.203 KiB)
Build problem: 78.574912 seconds (124.50 M allocations: 6.104 GiB, 4.43% gc time, 99.73% compilation time: 11% of which was recompilation)
KenCarp4(; linsolve = nothing, nlsolve = OrdinaryDiffEqNonlinearSolve.NLNewton{Rational{Int64}, Rational{Int64}, Rational{Int64}, Nothing}(1//100, 10, 1//5, 1//5, false, true, nothing), smooth_est = true, extrapolant = linear, step_limiter! = trivial_limiter!, autodiff = ADTypes.AutoForwardDiff(), concrete_jac = nothing,)stimstart = 30second
@unpack Istim, CaMKAct = sys
idxs = (t / 1000, CaMKAct)
sols = map((15, 30, 60, 90)) do dur
cb = build_stim_callbacks(Istim, stimstart + dur * second; period=1second, starttime=stimstart)
sol = solve(prob, alg; callback=cb)
end
plot(title="Simulation")
for (sol, dur, color) in zip(sols, (15, 30, 60, 90), (:blue, :red, :orange, :green))
plot!(sol, idxs=idxs, tspan=(0second, 205second), lab="$dur sec", color=color)
end
plot!(xlabel="Time (s)", ylabel="Active CaMKII fraction")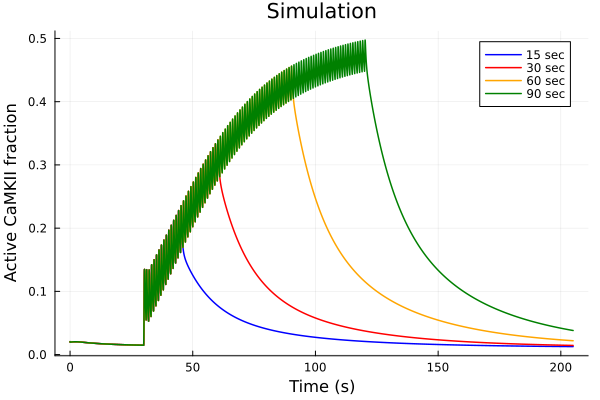
savefig("pacing-duration-sim.pdf")
savefig("pacing-duration-sim.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/pacing-duration-sim.png"Decay rates¶
Fit data from experiments and simulations against an exponential decay model.
Record 50 seconds after pacing ends.
ts = collect(range(0.0, stop=50.0, step=5.0)) ## in seconds
ydata_15 = fifteen[10:20]
ydata_30 = thirty[13:23]
ydata_60 = sixty[19:29]
ydata_90 = ninety[25:35]11-element Vector{Float64}:
13.89164293
13.73430494
13.47906779
13.37909541
13.29527631
13.14725458
13.13348484
13.06471802
13.08336721
12.99340616
13.04848118Simulation points
ysim_15 = sols[1](stimstart+15second:5second:stimstart+15second+50second ; idxs=sys.CaMKAct * 100).u
ysim_30 = sols[2](stimstart+30second:5second:stimstart+30second+50second ; idxs=sys.CaMKAct * 100).u
ysim_60 = sols[3](stimstart+60second:5second:stimstart+60second+50second ; idxs=sys.CaMKAct * 100).u
ysim_90 = sols[4](stimstart+90second:5second:stimstart+90second+50second ; idxs=sys.CaMKAct * 100).u11-element Vector{Float64}:
44.81684933831418
35.55473413472768
27.844712625927308
22.45556736675176
18.546953177388716
15.61258463816494
13.341815831583798
11.538576069044609
10.074965581307097
8.865462103033392
7.851377304622975Fit
fit_15 = solve(CurveFitProblem(ts, ydata_15), ExpSumFitAlgorithm(n=1, withconst=true))
fit_30 = solve(CurveFitProblem(ts, ydata_30), ExpSumFitAlgorithm(n=1, withconst=true))
fit_60 = solve(CurveFitProblem(ts, ydata_60), ExpSumFitAlgorithm(n=1, withconst=true))
fit_90 = solve(CurveFitProblem(ts, ydata_90), ExpSumFitAlgorithm(n=1, withconst=true))
fit_sim_15 = solve(CurveFitProblem(ts, ysim_15), ExpSumFitAlgorithm(n=1, withconst=true))
fit_sim_30 = solve(CurveFitProblem(ts, ysim_30), ExpSumFitAlgorithm(n=1, withconst=true))
fit_sim_60 = solve(CurveFitProblem(ts, ysim_60), ExpSumFitAlgorithm(n=1, withconst=true))
fit_sim_90 = solve(CurveFitProblem(ts, ysim_90), ExpSumFitAlgorithm(n=1, withconst=true))retcode: Success
alg: ExpSumFitAlgorithm
residuals mean: 2.4223047810003414e-16
u: [5.664165007372851, 39.10258079560866, -0.055270779777872675]Fitting results (experiments)¶
p1 = plot(ts, ydata_15, label="Exp 15 sec")
plot!(p1, ts, predict(fit_15), label="Fit", linestyle=:dash)
p2 = plot(ts, ydata_30, label="Exp 30 sec")
plot!(p2, ts, predict(fit_30), label="Fit", linestyle=:dash)
p3 = plot(ts, ydata_60, label="Exp 60 sec")
plot!(p3, ts, predict(fit_60), label="Fit", linestyle=:dash)
p4 = plot(ts, ydata_90, label="Exp 90 sec")
plot!(p4, ts, predict(fit_90), label="Fit", linestyle=:dash)
plot(p1, p2, p3, p4, layout=(2,2), xlabel="Time (s)", ylabel="CaMKII activity (AU)")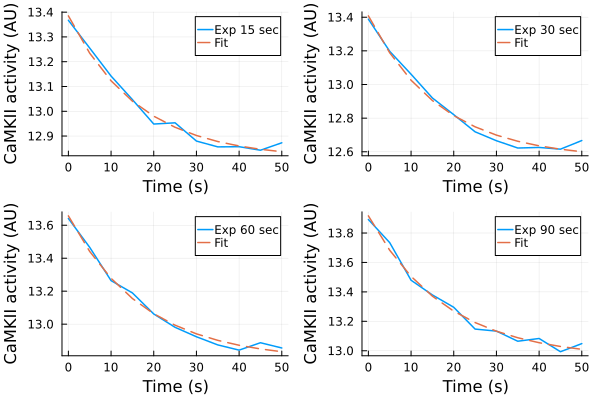
Fitting results (simulations)¶
p1s = plot(ts, ysim_15, label="Sim 15 sec")
plot!(p1s, ts, predict(fit_sim_15), label="Fit", linestyle=:dash)
p2s = plot(ts, ysim_30, label="Sim 30 sec")
plot!(p2s, ts, predict(fit_sim_30), label="Fit", linestyle=:dash)
p3s = plot(ts, ysim_60, label="Sim 60 sec")
plot!(p3s, ts, predict(fit_sim_60), label="Fit", linestyle=:dash)
p4s = plot(ts, ysim_90, label="Sim 90 sec")
plot!(p4s, ts, predict(fit_sim_90), label="Fit", linestyle=:dash)
plot(p1s, p2s, p3s, p4s, layout=(2,2), xlabel="Time (s)", ylabel="CaMKII activity (%)")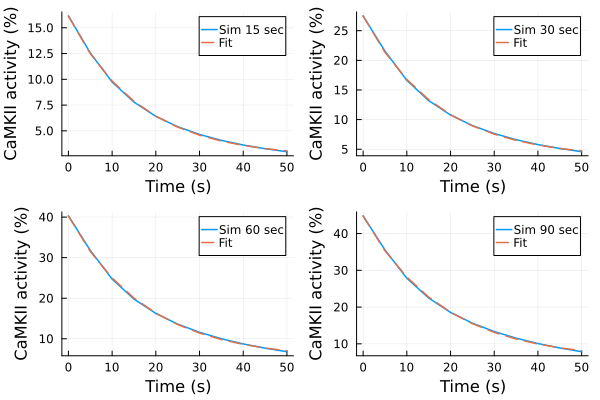
Decay time scales (tau)¶
tau_exp_15 = inv(-fit_15.u.λ[])
tau_exp_30 = inv(-fit_30.u.λ[])
tau_exp_60 = inv(-fit_60.u.λ[])
tau_exp_90 = inv(-fit_90.u.λ[])
tau_sim_15 = inv(-fit_sim_15.u.λ[])
tau_sim_30 = inv(-fit_sim_30.u.λ[])
tau_sim_60 = inv(-fit_sim_60.u.λ[])
tau_sim_90 = inv(-fit_sim_90.u.λ[])
println("The time scales for experiments: ")
for (tau, dur) in zip((tau_exp_15, tau_exp_30, tau_exp_60, tau_exp_90), (15, 30, 60, 90))
println("$dur sec pacing is $(round(tau; digits=2)) seconds.")
end
println("The time scales for simulations: ")
for (tau, dur) in zip((tau_sim_15, tau_sim_30, tau_sim_60, tau_sim_90), (15, 30, 60, 90))
println("$dur sec pacing is $(round(tau; digits=2)) seconds.")
endThe time scales for experiments:
15 sec pacing is 16.48 seconds.
30 sec pacing is 16.73 seconds.
60 sec pacing is 17.65 seconds.
90 sec pacing is 18.08 seconds.
The time scales for simulations:
15 sec pacing is 16.25 seconds.
30 sec pacing is 16.94 seconds.
60 sec pacing is 17.61 seconds.
90 sec pacing is 18.09 seconds.
Plot pacing time vs decay time scale
pacing_durations = [15.0, 30.0, 60.0, 90.0]
tau_experiments = [tau_exp_15, tau_exp_30, tau_exp_60, tau_exp_90]
tau_simulations = [tau_sim_15, tau_sim_30, tau_sim_60, tau_sim_90]
plot(pacing_durations, tau_experiments, label="Experiments", marker=:circle, color=:blue)
plot!(pacing_durations, tau_simulations, label="Simulations", marker=:square, color=:red)
plot!(title="Pacing Duration vs Decay Time Scale", xlabel="Pacing Duration (s)", ylabel="Decay Time Scale (s)")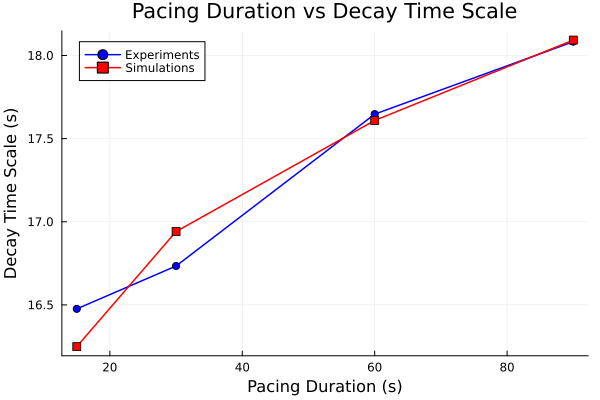
savefig("pacing-duration-tau.pdf")
savefig("pacing-duration-tau.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/pacing-duration-tau.png"Phosphorylated fraction¶
idxs = (sys.t / 1000, (sys.CaMKP + sys.CaMKA + sys.CaMKA2) * 100)
tspan = (0second, 205second)
plot(sols[1], idxs=idxs, tspan=tspan, lab="15 sec", color=:blue)
plot!(sols[2], idxs=idxs, tspan=tspan, lab="30 sec", color=:red)
plot!(sols[3], idxs=idxs, tspan=tspan, lab="60 sec", color=:orange)
plot!(sols[4], idxs=idxs, tspan=tspan, lab="90 sec", color=:green)
plot!(title="Simulation", xlabel="Time (s)", ylabel="Phosphorylated CaMKII (%)")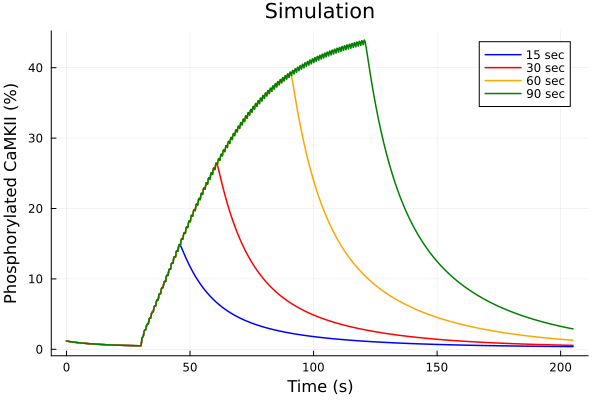
savefig("pacing-duration-phos.pdf")
savefig("pacing-duration-phos.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/pacing-duration-phos.png"This notebook was generated using Literate.jl.