1Hz vs 2Hz pacing
using Model
using Model: secondusing CSV
using CurveFit
using DataFrames
using DiffEqCallbacks
using DifferentialEquations
using ModelingToolkit
using OrdinaryDiffEqSDIRK
using Plots
using SteadyStateDiffEq
Plots.default(lw=1.5)Experiments¶
freqdf = CSV.read(joinpath(@__DIR__, "data/CaMKAR-freq.csv"), DataFrame)
ts = 0:5:205
onehz = freqdf[!, "1Hz (Mean)"]
onehz_error = freqdf[!, "1Hz (SD)"] ./ sqrt.(freqdf[!, "1Hz (N)"])
twohz = freqdf[!, "2Hz (Mean)"]
twohz_error = freqdf[!, "2Hz (SD)"] ./ sqrt.(freqdf[!, "2Hz (N)"])
plot(ts, onehz, yerr=onehz_error, lab="1 Hz", color=:blue, markerstrokecolor=:blue)
plot!(ts, twohz, yerr=twohz_error, lab="2 Hz", color=:red, markerstrokecolor=:red)
plot!(title="Experiment", xlabel="Time (s)", ylabel="CaMKII activity (AU)")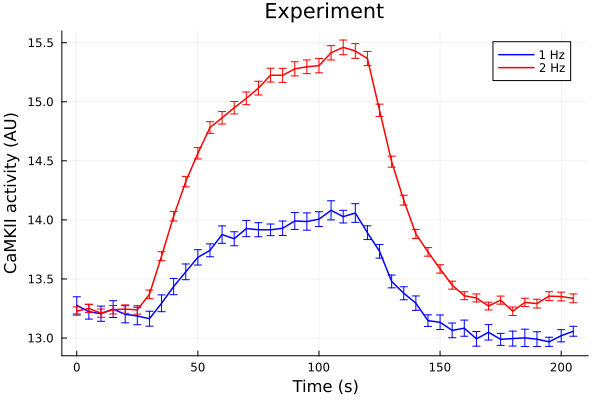
savefig("pacing-frequency-exp.pdf")
savefig("pacing-frequency-exp.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/pacing-frequency-exp.png"Simulations¶
@time "Build system" sys = Model.DEFAULT_SYS
tend = 205.0second
@time "Build problem" prob = ODEProblem(sys, [], tend)Build system: 0.000024 seconds (48 allocations: 2.203 KiB)
Build problem: 77.985386 seconds (124.30 M allocations: 6.094 GiB, 4.33% gc time, 99.71% compilation time: 55% of which was recompilation)
ODEProblem with uType Vector{Float64} and tType Float64. In-place: true
Initialization status: FULLY_DETERMINED
Non-trivial mass matrix: false
timespan: (0.0, 205000.0)
u0: 75-element Vector{Float64}:
830.0
830.0
0.0026
0.07192
0.07831
0.26081
0.00977
0.00188
0.09243
0.22156
⋮
0.12113
0.12113
0.12113
0.12113
0.12113
0.12113
150952.75035000002
13838.37602
-68.79268@unpack Istim = sys
stimstart = 30.0second
stimend = 120.0second
alg = KenCarp4()
callback = build_stim_callbacks(Istim, stimend; period=1second, starttime=stimstart)
@time "Solve problem" sol1 = solve(prob, alg; callback)
callback2 = build_stim_callbacks(Istim, stimend; period=0.5second, starttime=stimstart)
@time "Solve problem" sol2 = solve(prob, alg; callback=callback2)
idxs = (sys.t / 1000, sys.CaMKAct)
idxs_phos = (sys.t / 1000, (sys.CaMKP + sys.CaMKA + sys.CaMKA2))
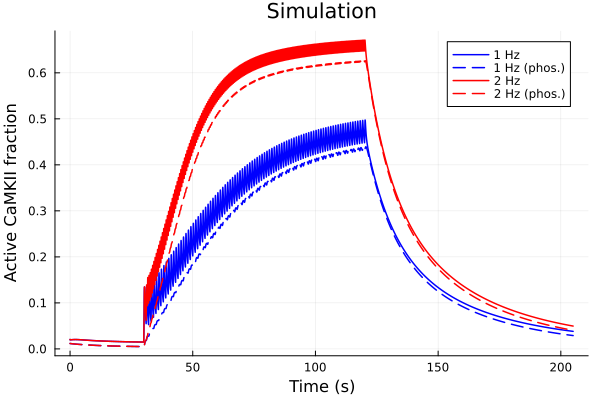
plot(sol1, idxs=idxs, lab="1 Hz", color=:blue)
plot!(sol1, idxs=idxs_phos, lab="1 Hz (phos.)", color=:blue, linestyle=:dash)
plot!(sol2, idxs=idxs, lab="2 Hz", color=:red)
plot!(sol2, idxs=idxs_phos, lab="2 Hz (phos.)", color=:red, linestyle=:dash)
plot!(title="Simulation", xlabel="Time (s)", ylabel="Active CaMKII fraction")Solve problem: 6.300281 seconds (14.51 M allocations: 668.325 MiB, 2.68% gc time, 83.85% compilation time)
Solve problem: 1.653427 seconds (74.49 k allocations: 17.323 MiB)
savefig("pacing-frequency-sim.pdf")
savefig("pacing-frequency-sim.png")"/home/github/actions-runner-2/_work/camkii-cardiomyocyte-model/camkii-cardiomyocyte-model/docs/pacing-frequency-sim.png"Decay rates¶
Fit against an exponential decay model.
Data from experiments: record 50 seconds after pacing ends
ts = collect(range(0.0, stop=50.0, step=5.0))
ydata_1hz = onehz[24:34]
ydata_2hz = twohz[24:34]
ysim_1hz = sol1(stimend:5second:stimend+50second; idxs=sys.CaMKAct).u
ysim_2hz = sol2(stimend:5second:stimend+50second; idxs=sys.CaMKAct).u
fit_1hz = solve(CurveFitProblem(ts, ydata_1hz), ExpSumFitAlgorithm(n=1, withconst=true))
fit_2hz = solve(CurveFitProblem(ts, ydata_2hz), ExpSumFitAlgorithm(n=1, withconst=true))
fit_1hz_sim = solve(CurveFitProblem(ts, ysim_1hz), ExpSumFitAlgorithm(n=1, withconst=true))
fit_2hz_sim = solve(CurveFitProblem(ts, ysim_2hz), ExpSumFitAlgorithm(n=1, withconst=true))retcode: Success
alg: ExpSumFitAlgorithm
residuals mean: 2.901719268906659e-17
u: [0.08537257813355881, 0.5564617133524954, -0.0599086652312363]Fitting results (experiments)
p1 = plot(ts, ydata_1hz, label="Exp 1 Hz")
plot!(p1, ts, predict(fit_1hz), label="Fit", linestyle=:dash)
p2 = plot(ts, ydata_2hz, label="Exp 2 Hz")
plot!(p2, ts, predict(fit_2hz), label="Fit", linestyle=:dash)
p3 = plot(ts, ysim_1hz, label="Sim 1 Hz")
plot!(p3, ts, predict(fit_1hz_sim), label="Fit", linestyle=:dash)
p4 = plot(ts, ysim_2hz, label="Sim 2 Hz")
plot!(p4, ts, predict(fit_2hz_sim), label="Fit", linestyle=:dash)
plot(p1, p2, p3, p4, layout=(2,2), xlabel="Time (s)", ylabel="CaMKII activity (AU)")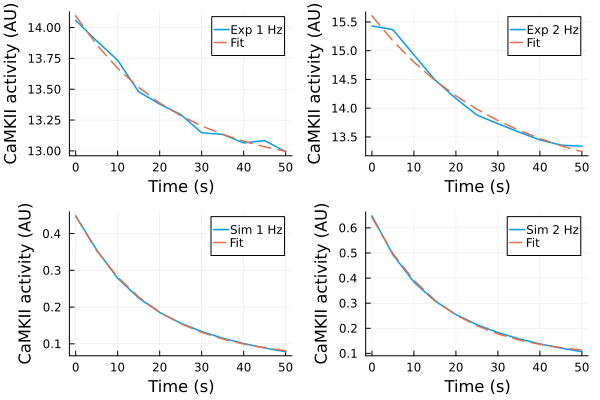
Decay time scales (tau) from fit parameters
tau_exp_1hz = inv(-fit_1hz.u.λ[])
tau_exp_2hz = inv(-fit_2hz.u.λ[])
tau_sim_1hz = inv(-fit_1hz_sim.u.λ[])
tau_sim_2hz = inv(-fit_2hz_sim.u.λ[])
println("The time scales for experiments: ")
for (tau, freq) in zip((tau_exp_1hz, tau_exp_2hz), (1, 2))
println("$freq Hz pacing is $(round(tau; digits=2)) seconds.")
end
println("The time scales for simulations: ")
for (tau, freq) in zip((tau_sim_1hz, tau_sim_2hz), (1, 2))
println("$freq Hz pacing is $(round(tau; digits=2)) seconds.")
endThe time scales for experiments:
1 Hz pacing is 24.31 seconds.
2 Hz pacing is 31.82 seconds.
The time scales for simulations:
1 Hz pacing is 18.09 seconds.
2 Hz pacing is 16.69 seconds.
This notebook was generated using Literate.jl.