using OrdinaryDiffEq
using OrdinaryDiffEqSDIRK
using SteadyStateDiffEq
using ModelingToolkit
using MitochondrialDynamics
using MitochondrialDynamics: second, μM, mV, mM, Hz, minute
import PythonPlot as plt
plt.matplotlib.rcParams["font.size"] = 14┌ Info: CondaPkg: Waiting for lock to be freed. You may delete this file if no other process is resolving.
└ lock_file = "/home/github/actions-runner-2/_work/mitodyn-ode/mitodyn-ode/.CondaPkg/lock"
14glc = 4.0:0.5:30.0
@time "Build system" @named sys = make_model()
@time "Build problem" prob = SteadyStateProblem(sys, [])Build system: 77.482052 seconds (122.19 M allocations: 6.024 GiB, 5.15% gc time, 99.94% compilation time: 28% of which was recompilation)
Build problem: 30.239062 seconds (60.48 M allocations: 2.995 GiB, 3.65% gc time, 99.77% compilation time: 13% of which was recompilation)
SteadyStateProblem with uType Vector{Float64}. In-place: true
u0: 9-element Vector{Float64}:
0.06
0.24
0.9
0.0087
0.0029
0.092
0.0002
0.001
0.057Change parameters
@unpack Glc, rETC, rHL, rF1, rPDH = sysModel sys:
Equations (9):
9 standard: see equations(sys)
Unknowns (9): see unknowns(sys)
x3(t)
x2(t)
AEC(t)
Pyr(t)
⋮
Parameters (65): see parameters(sys)
RestingCa
NCac
ActivatedCa
KatpCac
⋮
Observed (24): see observed(sys)Fig 6¶
prob_dm = SteadyStateProblem(sys, [rPDH=>0.5, rETC=>0.75, rHL=>1.4, rF1=>0.5])
prob_fccp = SteadyStateProblem(sys, [rHL=>5.0])
prob_rotenone = SteadyStateProblem(sys, [rETC=>0.1])
prob_oligomycin = SteadyStateProblem(sys, [rF1=>0.1])
prob_func_glc = (prob, ctx) -> remake(prob, p=[Glc => glc[ctx.sim_id]])#2 (generic function with 1 method)DM cells
alg = DynamicSS(KenCarp47())
trajectories = length(glc)
@time "Solve baseline" sols = map(glc) do g
_prob = remake(prob, p=[Glc => g])
solve(_prob, alg)
end
@time "Solve diabetic" solsDM = map(glc) do g
_prob = remake(prob_dm, p=[Glc => g])
solve(_prob, alg)
endSolve baseline: 16.964413 seconds (33.50 M allocations: 1.624 GiB, 2.47% gc time, 90.66% compilation time: 2% of which was recompilation)
Solve diabetic: 1.584379 seconds (6.09 M allocations: 291.932 MiB, 2.58% gc time, 6.91% compilation time)
function plot_fig6(sols, solsDM, glc; figsize=(10, 8), labels=["Baseline", "Diabetic"])
glc5 = glc ./ 5
numrows = 3
numcols = 3
fig, ax = plt.subplots(numrows, numcols; figsize)
@unpack G3P = sys
ax[0, 0].plot(glc5, extract(sols, G3P * 1000), label=labels[1])
ax[0, 0].plot(glc5, extract(solsDM, G3P * 1000), label=labels[2])
ax[0, 0].set_title("j", loc="left")
ax[0, 0].set(ylabel="G3P (μM)")
@unpack Pyr = sys
ax[0, 1].plot(glc5, extract(sols, Pyr * 1000), label=labels[1])
ax[0, 1].plot(glc5, extract(solsDM, Pyr * 1000), label=labels[2])
ax[0, 1].set_title("k", loc="left")
ax[0, 1].set(ylabel="Pyruvate (μM)")
@unpack NADH_c, NAD_c = sys
ax[0, 2].plot(glc5, extract(sols, NADH_c/NAD_c), label=labels[1])
ax[0, 2].plot(glc5, extract(solsDM, NADH_c/NAD_c), label=labels[2])
ax[0, 2].set_title("l", loc="left")
ax[0, 2].set(ylabel="Cyto. NADH:NAD (ratio)")
@unpack NADH_m, NAD_m = sys
ax[1, 0].plot(glc5, extract(sols, NADH_m/NAD_m), label=labels[1])
ax[1, 0].plot(glc5, extract(solsDM, NADH_m/NAD_m), label=labels[2])
ax[1, 0].set_title("m", loc="left")
ax[1, 0].set(ylabel="Mito. NADH:NAD (ratio)")
@unpack Ca_c = sys
ax[1, 1].plot(glc5, extract(sols, Ca_c * 1000), label=labels[1])
ax[1, 1].plot(glc5, extract(solsDM, Ca_c * 1000), label=labels[2])
ax[1, 1].set_title("n", loc="left")
ax[1, 1].set(ylabel="Cyto. calcium (μM)")
@unpack Ca_m = sys
ax[1, 2].plot(glc5, extract(sols, Ca_m * 1000), label=labels[1])
ax[1, 2].plot(glc5, extract(solsDM, Ca_m * 1000), label=labels[2])
ax[1, 2].set_title("o", loc="left")
ax[1, 2].set(ylabel="Mito. calcium (μM)")
@unpack ΔΨm = sys
ax[2, 0].plot(glc5, extract(sols, ΔΨm * 1000), label=labels[1])
ax[2, 0].plot(glc5, extract(solsDM, ΔΨm * 1000), label=labels[2])
ax[2, 0].set_title("p", loc="left")
ax[2, 0].set(xlabel="Glucose (X)", ylabel="ΔΨ (mV)")
@unpack ATP_c, ADP_c = sys
ax[2, 1].plot(glc5, extract(sols, ATP_c/ADP_c), label=labels[1])
ax[2, 1].plot(glc5, extract(solsDM, ATP_c/ADP_c), label=labels[2])
ax[2, 1].set_title("q", loc="left")
ax[2, 1].set(xlabel="Glucose (X)", ylabel="ATP:ADP (ratio)")
@unpack degavg = sys
ax[2, 2].plot(glc5, extract(sols, degavg), label=labels[1])
ax[2, 2].plot(glc5, extract(solsDM, degavg), label=labels[2])
ax[2, 2].set_title("r", loc="left")
ax[2, 2].set(xlabel="Glucose (X)", ylabel="Avg. node degree (ratio)")
for i in 0:numrows-1, j in 0:numcols-1
ax[i, j].grid()
ax[i, j].legend()
end
fig.tight_layout()
return fig
endplot_fig6 (generic function with 1 method)fig6 = plot_fig6(sols, solsDM, glc)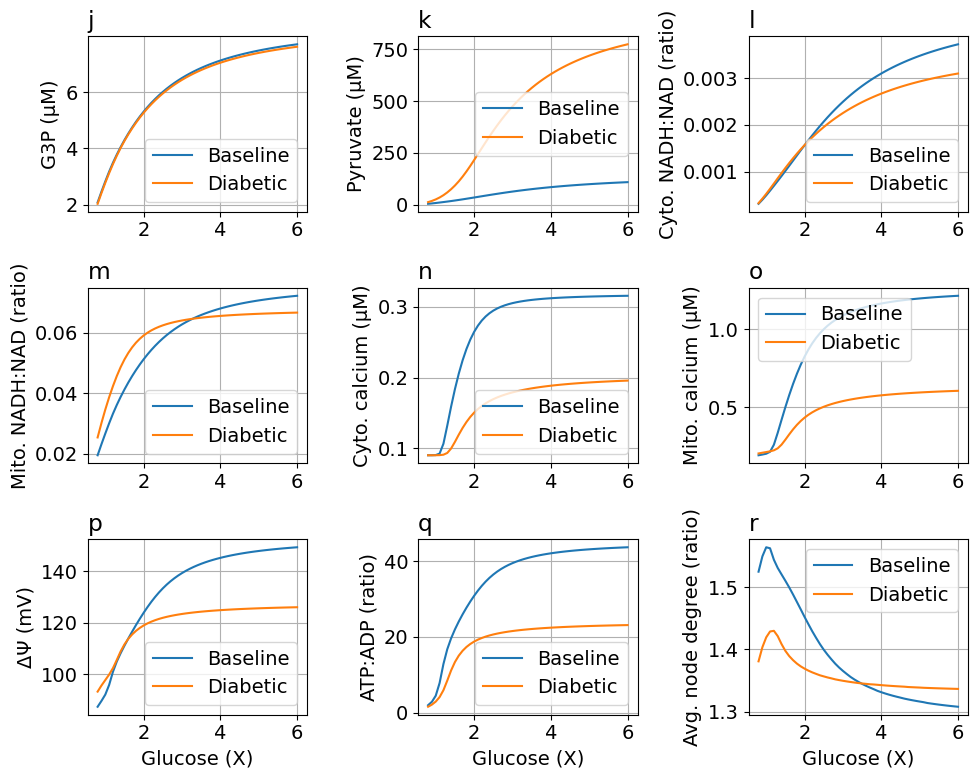
Export figure
exportTIF(fig6, "Fig7-DM-steadystates.tif")Python: NoneFigure 7¶
@time "Solve baseline" sols = map(glc) do g
_prob = remake(prob, p=[Glc => g])
solve(_prob, alg)
end
@time "Solve diabetic" solsDM = map(glc) do g
_prob = remake(prob_dm, p=[Glc => g])
solve(_prob, alg)
end
@time "Solve FCCP" solsFCCP = map(glc) do g
_prob = remake(prob_fccp, p=[Glc => g])
solve(_prob, alg)
end
@time "Solve Rotenone" solsRot = map(glc) do g
_prob = remake(prob_oligomycin, p=[Glc => g])
solve(_prob, alg)
end
@time "Solve Oligomycin" solsOligo = map(glc) do g
_prob = remake(prob_rotenone, p=[Glc => g])
solve(_prob, alg)
endSolve baseline: 1.365219 seconds (6.10 M allocations: 292.167 MiB, 7.11% gc time, 6.70% compilation time)
Solve diabetic: 1.318408 seconds (6.09 M allocations: 291.842 MiB, 7.94% gc time, 6.60% compilation time)
Solve FCCP: 1.584639 seconds (6.09 M allocations: 291.857 MiB, 23.33% gc time, 5.48% compilation time)
Solve Rotenone: 1.299282 seconds (6.09 M allocations: 291.735 MiB, 3.49% gc time, 6.78% compilation time)
Solve Oligomycin: 1.307987 seconds (6.09 M allocations: 292.123 MiB, 5.68% gc time, 6.71% compilation time)
function plot_fig7(sols, solsDM, solsFCCP, solsRot, solsOligo, glc; figsize=(12, 6))
sys = sols[begin].prob.f.sys
@unpack J_HL, J_ANT = sys
# Gather ATP synthesis rate (fusion) and proton leak rate (fission)
jHL_baseline = extract(sols, J_HL)
jANT_baseline = extract(sols, J_ANT)
ff_baseline = jANT_baseline ./ jHL_baseline
jHL_dm = extract(solsDM, J_HL)
jANT_dm = extract(solsDM, J_ANT)
ff_dm = jANT_dm ./ jHL_dm
jHL_fccp = extract(solsFCCP, J_HL)
jANT_fccp = extract(solsFCCP, J_ANT)
ff_fccp = jANT_fccp ./ jHL_fccp
jHL_rot = extract(solsRot, J_HL)
jANT_rot = extract(solsRot, J_ANT)
ff_rot = jANT_rot ./ jHL_rot
jHL_oligo = extract(solsOligo, J_HL)
jANT_oligo = extract(solsOligo, J_ANT)
ff_oligo = jANT_oligo ./ jHL_oligo
glc5 = glc ./ 5
fig, ax = plt.subplots(1, 2; figsize)
ax[0].plot(glc5, ff_baseline, "b-", label="Baseline")
ax[0].plot(glc5, ff_dm, "r--", label="Diabetic")
ax[0].plot(glc5, ff_rot, "g--", label="Rotenone")
ax[0].plot(glc5, ff_oligo, "c--", label="Oligomycin")
ax[0].plot(glc5, ff_fccp, "k--", label="Uncoupler")
ax[0].set(xlabel="Glucose (X)", ylabel="Fusion rate / Fission rate (ratio)", xlim=(0.0, 6.0), ylim=(0.0, 2.5))
ax[0].set_title("a", loc="left")
ax[0].grid()
ax[0].legend()
ax[1].plot(jHL_baseline, jANT_baseline, "bo-", label="Baseline")
ax[1].plot(jHL_dm, jANT_dm, "ro-", label="Diabetic")
ax[1].plot(jHL_rot, jANT_rot, "go-", label="Rotenone")
ax[1].plot(jHL_oligo, jANT_oligo, "co-", label="Oligomycin")
ax[1].plot(jHL_fccp, jANT_fccp, "ko-", label="Uncoupler")
ax[1].set(xlabel="Proton leak rate (mM/s)", ylabel="ATP synthase rate (mM/s)", xlim=(0.0, 0.45), ylim=(0.0, 0.15))
ax[1].set_title("b", loc="left")
ax[1].grid()
ax[1].legend()
fig.tight_layout()
return fig
endplot_fig7 (generic function with 1 method)fig7 = plot_fig7(sols, solsDM, solsFCCP, solsRot, solsOligo, glc)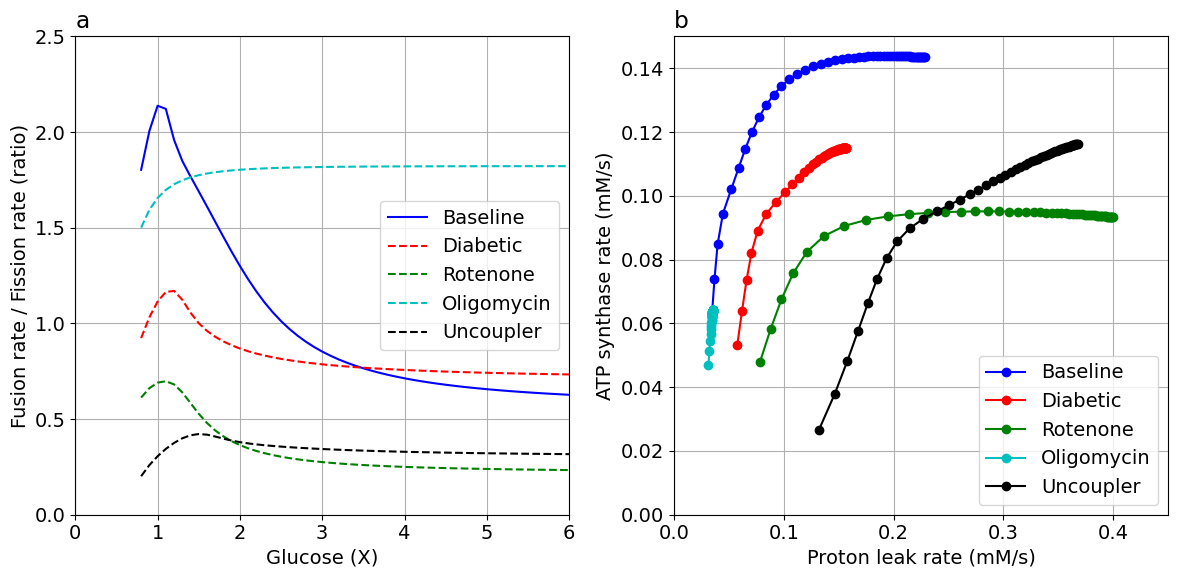
Export figure
exportTIF(fig7, "Fig8.tif")Python: NoneThis notebook was generated using Literate.jl.