using OrdinaryDiffEq
using OrdinaryDiffEqSDIRK
using ModelingToolkit
using MitochondrialDynamics
using MitochondrialDynamics: second, μM, mV, mM, Hz, minute, hil
import PythonPlot as plt
plt.matplotlib.rcParams["font.size"] = 14┌ Info: CondaPkg: Waiting for lock to be freed. You may delete this file if no other process is resolving.
└ lock_file = "/home/github/actions-runner-2/_work/mitodyn-ode/mitodyn-ode/.CondaPkg/lock"
14@time "Build system" @named sys = make_model()
@unpack Ca_c, Glc, kATPCa, kATP = sys
alg = TRBDF2()Build system: 78.463161 seconds (122.19 M allocations: 6.024 GiB, 4.88% gc time, 99.94% compilation time: 28% of which was recompilation)
TRBDF2(; linsolve = nothing, nlsolve = OrdinaryDiffEqNonlinearSolve.NLNewton{Rational{Int64}, Rational{Int64}, Rational{Int64}, Nothing}(1//100, 10, 1//5, 1//5, false, true, nothing), smooth_est = true, extrapolant = linear, step_limiter! = trivial_limiter!, autodiff = ADTypes.AutoForwardDiff(), concrete_jac = nothing,)Calcium oscillation function
function cac_wave(; ca_base = 0.09μM, ca_act = 0.25μM, n=4, katp=25, amplitude=0.5, period=2minute)
@variables t Ca_c(t) ATP_c(t) ADP_c(t)
@parameters (RestingCa=ca_base, ActivatedCa=ca_act, NCac=n, KatpCac=katp)
x = 5 * ((t / period) % 1.0) ## An oscillating function
w = (x * exp(1 - x))^4 ## Scale from 0 to 1
caceq = Ca_c ~ RestingCa + ActivatedCa * hil(ATP_c, KatpCac * ADP_c, NCac) * (1 + amplitude * (2w-1))
return caceq
end
@time "Build oscillating system" @named sysosci = make_model(; caceq=cac_wave(amplitude=0.8))Build oscillating system: 1.022561 seconds (2.27 M allocations: 113.931 MiB, 98.14% compilation time)
Model sysosci:
Equations (9):
9 standard: see equations(sysosci)
Unknowns (9): see unknowns(sysosci)
x3(t)
x2(t)
AEC(t)
Pyr(t)
⋮
Parameters (65): see parameters(sysosci)
RestingCa
NCac
ActivatedCa
KatpCac
⋮
Observed (24): see observed(sysosci)tend = 4000.0
ts = range(tend-480, tend; length=201)
@time "Build oscillating problem" prob = ODEProblem(sysosci, [], tend, [Glc => 10mM])
@time "Solve oscillating problem" sol = solve(prob, alg, saveat=ts)┌ Warning: `SciMLBase.ODEProblem(sys, u0, tspan, p; kw...)` is deprecated. Use
│ `SciMLBase.ODEProblem(sys, merge(if isempty(u0)
│ Dict()
│ else
│ Dict(unknowns(sys) .=> u0)
│ end, Dict(p)), tspan)` instead.
└ @ ModelingToolkitBase ~/.julia/packages/ModelingToolkitBase/eSs1G/src/deprecations.jl:53
Build oscillating problem: 29.562810 seconds (59.23 M allocations: 2.922 GiB, 3.65% gc time, 99.56% compilation time: 41% of which was recompilation)
Solve oscillating problem: 4.729118 seconds (9.58 M allocations: 480.030 MiB, 2.54% gc time, 99.40% compilation time)
retcode: Success
Interpolation: 1st order linear
t: 201-element Vector{Float64}:
3520.0
3522.4
3524.8
3527.2
3529.6
3532.0
3534.4
3536.8
3539.2
3541.6
⋮
3980.8
3983.2
3985.6
3988.0
3990.4
3992.8
3995.2
3997.6
4000.0
u: 201-element Vector{Vector{Float64}}:
[0.03089399783100657, 0.21294490286270895, 0.9825783019375932, 0.034741355734069966, 0.005308721702614443, 0.12216210667236822, 0.0008340842726265699, 0.0031095823201684834, 0.10663461645894881]
[0.030979340677080965, 0.21323152736447887, 0.9835702897053655, 0.034922414202243804, 0.005313862543507346, 0.12421422163017364, 0.0008280421786997974, 0.0031207249257225083, 0.10715081915928604]
[0.03104328641535978, 0.2134408656067794, 0.9845599426306899, 0.03514545703291697, 0.0053195204531972145, 0.126541107921258, 0.000818851350805462, 0.0031339543841007907, 0.10777071593357301]
[0.031085424884643054, 0.21357208922059154, 0.9855255183028525, 0.03541123416780667, 0.005325599352460509, 0.12910953333957434, 0.0008064122858256864, 0.0031492405181024444, 0.10849291969251386]
[0.03110709511146832, 0.21363115978315583, 0.986377208513856, 0.03571386987850111, 0.005331539405072436, 0.1316941652087737, 0.0007907872452649076, 0.0031658562663478937, 0.1092826221410523]
[0.031110612318601977, 0.21362692593825008, 0.9870684368068259, 0.03604272105169195, 0.005336914428653108, 0.13410716744114734, 0.0007722817633589922, 0.0031828881062293036, 0.11009605572448464]
[0.031097574397663173, 0.21356509367834076, 0.9876127766815963, 0.03638943714493726, 0.005341644201114228, 0.13629691849256018, 0.000751195761070688, 0.003199788661942486, 0.11090700584969838]
[0.031069718259257785, 0.21345185667026087, 0.9880252466856686, 0.036744841591319174, 0.005345647110911994, 0.1382099417170874, 0.0007278766404424178, 0.0032159721448967423, 0.11168737227769793]
[0.03103167582963861, 0.21330353117543804, 0.988349451377193, 0.03708313997276788, 0.005348860426172982, 0.13977956086590823, 0.0007037340645244035, 0.003230213488688383, 0.1123776702378804]
[0.030985826474912, 0.21312844874209622, 0.988601804420612, 0.037393385517981034, 0.005351363784746499, 0.14103290881916777, 0.0006795925551900223, 0.0032423127465649273, 0.11296872952184253]
⋮
[0.029699941155191978, 0.2086434227740685, 0.9790982205169241, 0.03502507536513055, 0.00529435773066293, 0.11685744609140136, 0.0007583988460648041, 0.0030928907348183807, 0.10600309056613003]
[0.029869607368167937, 0.2092805981586058, 0.9787166462047864, 0.034728856357758534, 0.005292817681652869, 0.11629019488985823, 0.0007841417587196497, 0.003086386141450851, 0.10567140927990361]
[0.030042282752861898, 0.2099203986281559, 0.9786701240389666, 0.03455061547372823, 0.005292567717733687, 0.11618706866164653, 0.0008034979187916833, 0.0030834073244946593, 0.10550868271973231]
[0.03021249022577004, 0.21054254949004655, 0.9788875765182803, 0.03445687644866898, 0.005293267731771012, 0.11642086323241693, 0.0008177068509192435, 0.0030829164534722556, 0.1054663238268995]
[0.030375058653252685, 0.2111292861763571, 0.9793211324139377, 0.03442286398681484, 0.005294839282753891, 0.11697627569686751, 0.0008277705984010609, 0.003084525729788424, 0.10552362920690464]
[0.030526991402887598, 0.21167070371354985, 0.9799363360715191, 0.034437951408963434, 0.005297206466122967, 0.11782764648586917, 0.0008342083553948909, 0.0030880320568332817, 0.10566949622611392]
[0.030666739133556803, 0.21216188598485589, 0.9807144330996417, 0.03449860228735233, 0.005300336947505769, 0.11897095709645339, 0.0008372331799547996, 0.003093364086440303, 0.10590143134460775]
[0.030791697565591303, 0.2125947098129923, 0.9816132698839043, 0.03460045541594219, 0.005304198283806756, 0.12041586808662859, 0.0008371490036792366, 0.0031005212391325586, 0.10622156031175777]
[0.03089824008574765, 0.2129581536438988, 0.9825775631836174, 0.03474045537802495, 0.005308748243926601, 0.12217473099879717, 0.0008342120982540428, 0.0031096115142136313, 0.1066362631191568]function plot_fig5(sol, figsize=(10, 10))
ts = sol.t
tsm = ts ./ 60
@unpack Ca_c, Ca_m, ATP_c, ADP_c, ΔΨm, degavg, J_ANT, J_HL = sys
fig, ax = plt.subplots(5, 1; figsize)
ax[0].plot(tsm, sol[Ca_c * 1000], label="Cyto")
ax[0].plot(tsm, sol[Ca_m * 1000], label="Mito")
ax[0].set_title("a", loc="left")
ax[0].set_ylabel("Calcium (μM)", fontsize=12)
ax[0].legend(loc="center left")
ax[1].plot(tsm, sol[ATP_c / ADP_c])
ax[1].set_title("b", loc="left")
ax[1].set_ylabel("ATP:ADP (ratio)", fontsize=12)
ax[2].plot(tsm, sol[ΔΨm * 1000])
ax[2].set_title("c", loc="left")
ax[2].set_ylabel("ΔΨm (mV)", fontsize=12)
ax[3].plot(tsm, sol[degavg], label="Average node degree")
ax[3].set_title("d", loc="left")
ax[3].set_ylabel("a.u.")
ax[3].legend(loc="center left")
ax[4].plot(tsm, sol[J_ANT], label="ATP export")
ax[4].plot(tsm, sol[J_HL], label="H leak")
ax[4].set_title("e", loc="left")
ax[4].set_ylabel("Rate (mM/s)")
ax[4].set(xlabel="Time (minute)")
ax[4].legend(loc="center left")
for i in 0:4
ax[i].grid()
ax[i].set_xlim(tsm[begin], tsm[end])
end
plt.tight_layout()
return fig
endplot_fig5 (generic function with 2 methods)fig5 = plot_fig5(sol)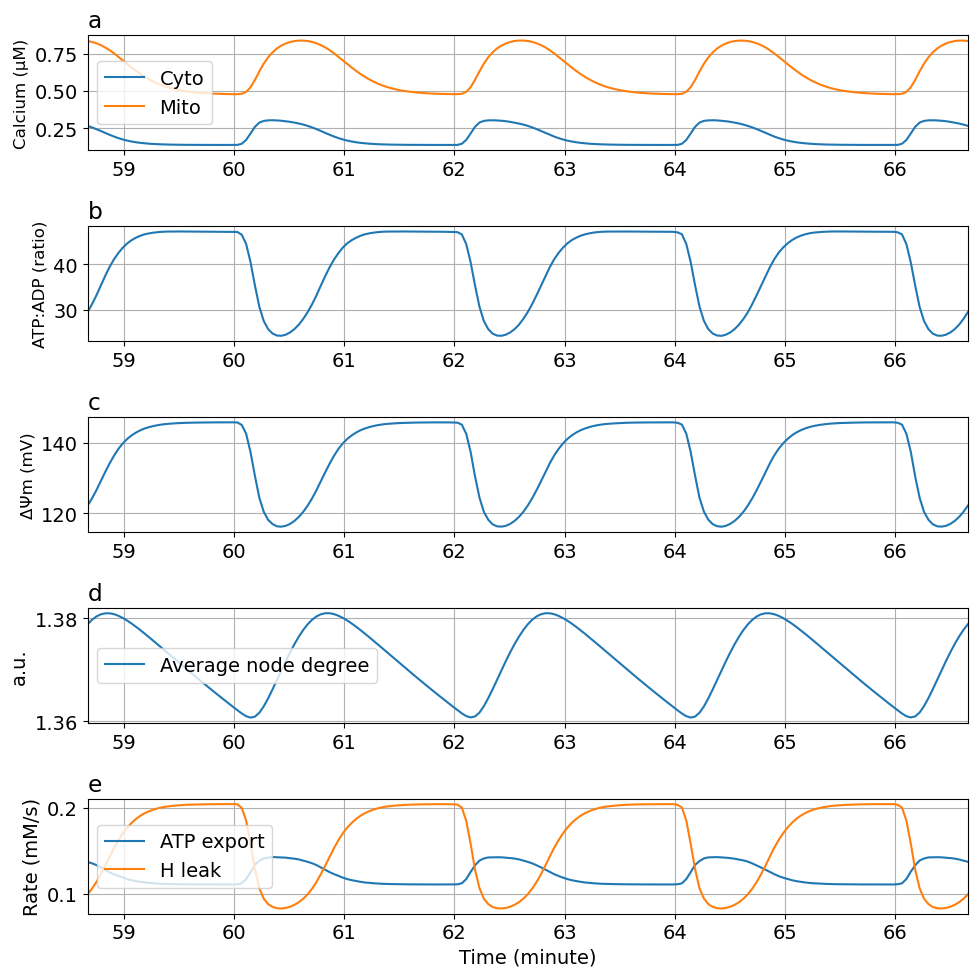
Export figure
exportTIF(fig5, "Fig6-ca-oscillation.tif")Python: NoneTuning ca-dependent ATP consumption rate (kATPCa) kATPCa : 90 -> 10
prob2 = ODEProblem(sysosci, [], tend, [Glc => 10mM, kATPCa=>10Hz/mM, kATP=>0.055Hz])
@time "Solve tuned problem 2" sol2 = solve(prob2, alg, saveat=ts)
plot_fig5(sol2)┌ Warning: `SciMLBase.ODEProblem(sys, u0, tspan, p; kw...)` is deprecated. Use
│ `SciMLBase.ODEProblem(sys, merge(if isempty(u0)
│ Dict()
│ else
│ Dict(unknowns(sys) .=> u0)
│ end, Dict(p)), tspan)` instead.
└ @ ModelingToolkitBase ~/.julia/packages/ModelingToolkitBase/eSs1G/src/deprecations.jl:53
Solve tuned problem 2: 0.004835 seconds (1.82 k allocations: 95.336 KiB)
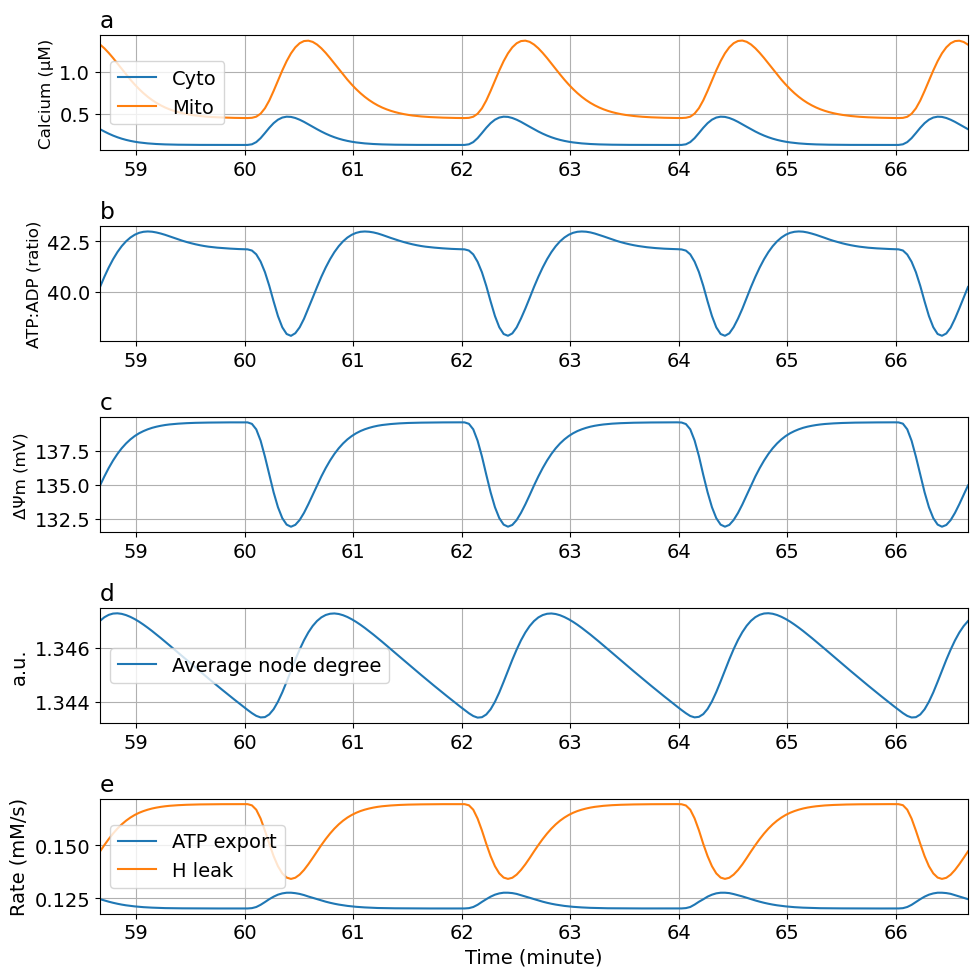
kATPCa : 90 -> 0.1
prob4 = ODEProblem(sysosci, [], tend, [Glc => 10mM, kATPCa=>0.1Hz/mM, kATP=>0.06Hz])
@time "Solve tuned problem 4" sol4 = solve(prob4, alg, saveat=ts)
plot_fig5(sol4)┌ Warning: `SciMLBase.ODEProblem(sys, u0, tspan, p; kw...)` is deprecated. Use
│ `SciMLBase.ODEProblem(sys, merge(if isempty(u0)
│ Dict()
│ else
│ Dict(unknowns(sys) .=> u0)
│ end, Dict(p)), tspan)` instead.
└ @ ModelingToolkitBase ~/.julia/packages/ModelingToolkitBase/eSs1G/src/deprecations.jl:53
Solve tuned problem 4: 0.004589 seconds (1.82 k allocations: 95.289 KiB)
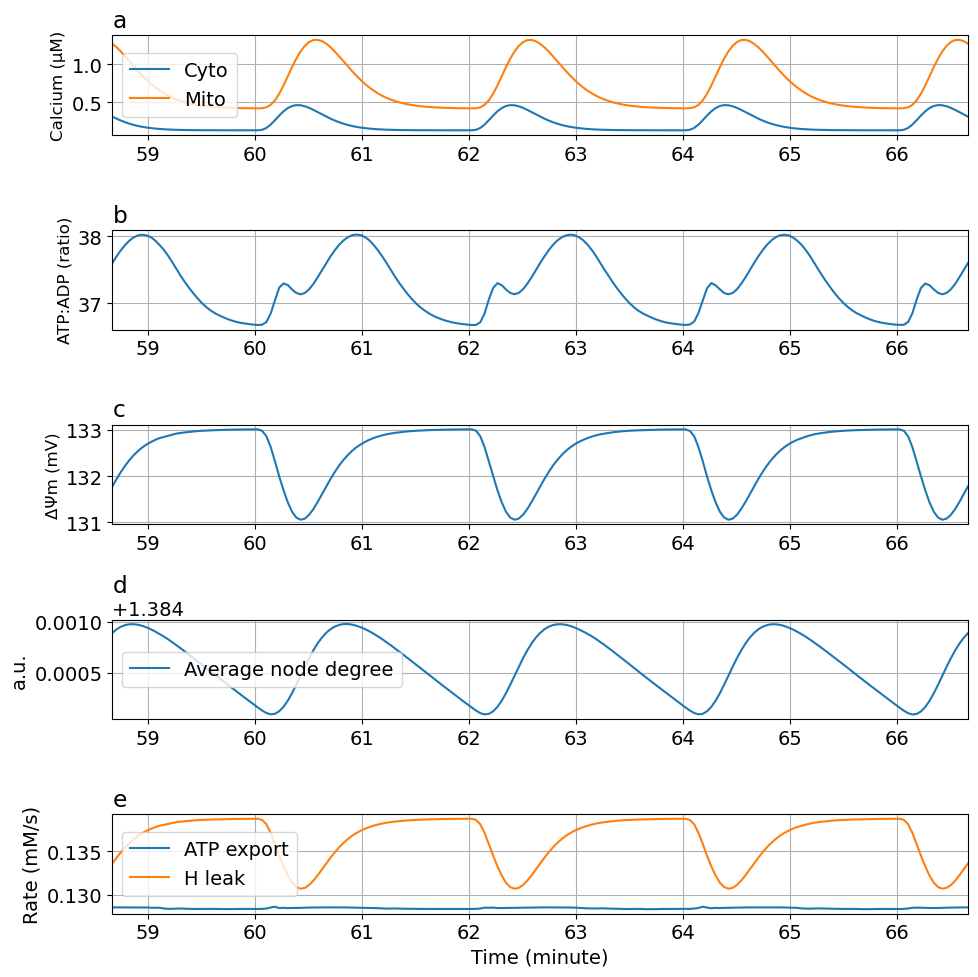
This notebook was generated using Literate.jl.