Fig 6.3¶
Two component pathway
using OrdinaryDiffEq
using Catalyst
using Plots
@time "Build system" rn603 = @reaction_network begin
@discrete_events begin
(t == 1.0) => [L => 3.0]
(t == 3.0) => [L => 0.0]
end
(k1 * L, km1), R <--> RL
k2, RL + P --> RL + Ps
k3, Ps --> P
endBuild system: 18.396671 seconds (39.55 M allocations: 1.938 GiB, 2.06% gc time, 99.96% compilation time: 9% of which was recompilation)
Loading...
up603 = Dict(:k1 => 5.0, :km1 => 1.0, :k2 => 6.0, :k3 => 2.0, :L => 0.0, :R => 3.0, :RL => 0.0, :P => 8.0, :Ps => 0.0)
tend = 10.0
@time "Build problem" prob603 = ODEProblem(rn603, up603, (0.0, tend); remove_conserved=true)
@time "Solve problem" sol603 = solve(prob603, TRBDF2(); tstops=[1.0, 3.0])
plot(sol603, idxs=[:RL, :Ps, :L], labels=["RL" "P*" "L"], title="Fig. 6.3 (A)", xlabel="Time", ylabel="Concentration")Build problem: 48.825502 seconds (92.04 M allocations: 4.533 GiB, 1.36% gc time, 99.85% compilation time: 29% of which was recompilation)
Solve problem: 8.787532 seconds (21.15 M allocations: 1.073 GiB, 1.36% gc time, 99.81% compilation time)
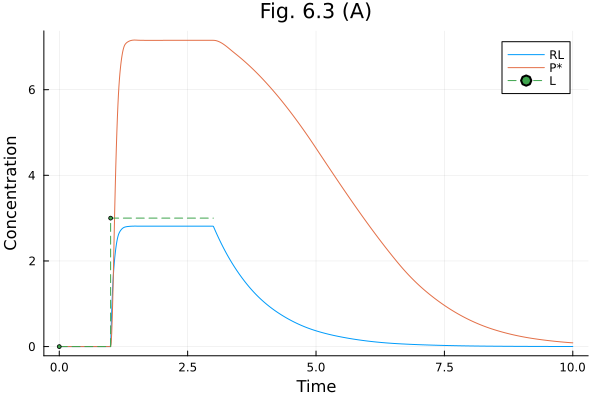
Fig 6.5¶
Model of G-protein signalling pathway
using OrdinaryDiffEq
using SteadyStateDiffEq
using Catalyst
using Plots
@time "Build system" rn605 = @reaction_network begin
@discrete_events begin
(t == 200.0) => [L => 1e-9]
(t == 800.0) => [L => 0.0]
end
kRL * L, R --> RL
kRLm, RL --> R
kGa, RL + G --> RL + Ga + Gbg
kGd0, Ga --> Gd
kG1, Gd + Gbg --> G
endBuild system: 0.011879 seconds (9.41 k allocations: 448.250 KiB, 89.64% compilation time)
Loading...
tend = 1200.0
up605 = Dict(:R => 4e3, :RL => 0.0, :G => 1e4, :Ga => 0.0, :Gd => 0.0, :Gbg => 0.0, :kRL => 2e6, :kRLm => 0.01, :kGa => 1e-5, :kGd0 => 0.11, :kG1 => 1.0, :L => 0.0)
@time "Build problem" prob605 = ODEProblem(rn605, up605, (0.0, tend); remove_conserved=true)
@time "Solve problem" sol = solve(prob605, KenCarp47(); tstops=[200.0, 800.0])
plot(sol, idxs=[:RL, :Ga], labels=["RL" "Ga"], title="Fig. 6.05 (A)", xlabel="Time", ylabel="Concentration")Build problem: 3.957394 seconds (7.51 M allocations: 395.214 MiB, 1.14% gc time, 98.97% compilation time: 18% of which was recompilation)
Solve problem: 9.237056 seconds (20.36 M allocations: 1.017 GiB, 1.09% gc time, 99.83% compilation time)
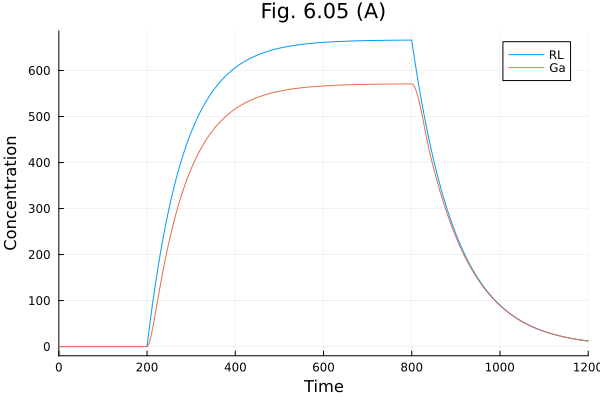
Fig 6.5 B¶
lrange = range(0, 20 * 1e-9, length=101)
@time "Build system" rn605b = @reaction_network begin
kRL * L, R --> RL
kRLm, RL --> R
kGa, RL + G --> RL + Ga + Gbg
kGd0, Ga --> Gd
kG1, Gd + Gbg --> G
end
@time "Build problem" prob605b = SteadyStateProblem(rn605b, up605; remove_conserved=true)
@time "Ensemble simulations" sols605 = map(lrange) do lval
_p = remake(prob605b; p=[:L => lval])
sol = solve(_p, DynamicSS(KenCarp47()))
end
ga = [sol[:Ga] for sol in sols605]
rl = [sol[:RL] for sol in sols605]
plot(lrange .* 1e9, [ga , rl], label=["Ga" "RL"], xlabel="L (nM)", ylabel="Steady-state abundance", title = "Fig. 6.5 (B)")Build system: 0.559754 seconds (609.09 k allocations: 29.379 MiB, 99.82% compilation time)
Build problem: 4.325998 seconds (10.89 M allocations: 559.042 MiB, 1.45% gc time, 99.37% compilation time)
Ensemble simulations: 10.982279 seconds (25.64 M allocations: 1.260 GiB, 1.06% gc time, 92.76% compilation time)
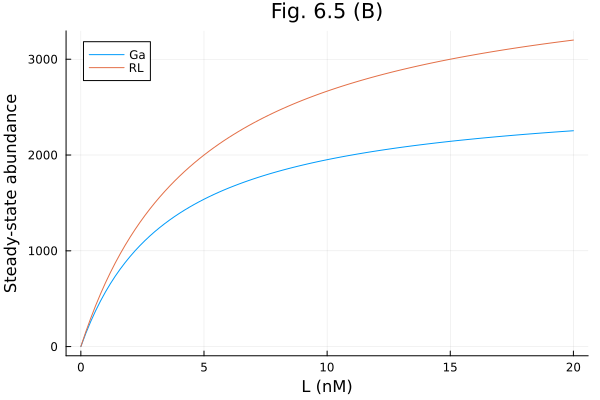
Fig 6.07¶
Goldbeter Koshland switch
using Plots
f607(w, K1, K2) = w * (1 - w + K1)/((1 - w) * (w + K2))
yy = 0:0.001:0.999
xx1 = f607.(yy, 1, 1)
xx2 = f607.(yy, 0.1, 0.1)
xx3 = f607.(yy, 0.01, 0.01)
plot(xx1, yy, label="K1=K2=1", xlabel="Stimulus", ylabel="Response", title="Fig 6.07", xlims=(0, 3), ylims=(0, 1))
plot!(xx2, yy, label="K1=K2=0.1")
plot!(xx3, yy, label="K1=K2=0.01")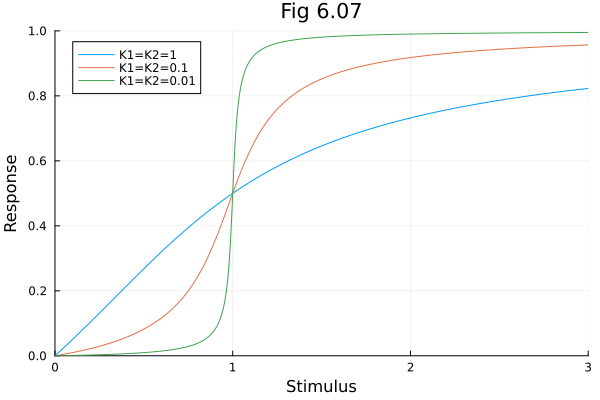
Fig 6.14¶
Model of E. coli chemotaxis signalling pathway
using OrdinaryDiffEq
using Catalyst
using Plots
@time "Build system" rn614 = @reaction_network begin
@discrete_events begin
(t == 10.0) => [L => 40.0]
(t == 30.0) => [L => 80.0]
end
mm(Am, k1 * BP, KM1), Am => A
mm(AmL, k2 * BP, KM2), AmL => AL
km1 * R , A => Am
km2 * R , AL => AmL
(k3 * L, km3), Am <--> AmL
(k4 * L, km4), A <--> AL
(k5 * Am, km5), B <--> BP
endBuild system: 0.126191 seconds (325.90 k allocations: 16.193 MiB, 41.47% gc time, 99.02% compilation time)
Loading...
up614 = Dict(
:Am => 0.0360,
:AmL => 1.5593,
:A => 0.0595,
:AL => 0.3504,
:B => 0.7356,
:BP => 0.2644,
:k1 => 200.0,
:k2 => 1.0,
:k3 => 1.0,
:km1 => 1.0,
:km2 => 1.0,
:km3 => 1.0,
:k4 => 1.0,
:km4 => 1.0,
:k5 => 0.05,
:km5 => 0.005,
:KM1 => 1.0,
:KM2 => 1.0,
:L => 20.0,
:R => 1.0,
)
tend = 50.0
@time "Build problem" prob614 = ODEProblem(rn614, up614, (0.0, tend); remove_conserved=true)
@time "Solve problem" sol614 = solve(prob614, FBDF(); tstops=[10.0, 30.0])
plot(sol614, idxs=[:Am], title="Fig 6.14", xlabel="Time", ylabel="Concentration")Build problem: 1.765099 seconds (2.31 M allocations: 121.739 MiB, 97.50% compilation time)
Solve problem: 5.464921 seconds (12.63 M allocations: 663.385 MiB, 4.82% gc time, 99.72% compilation time)
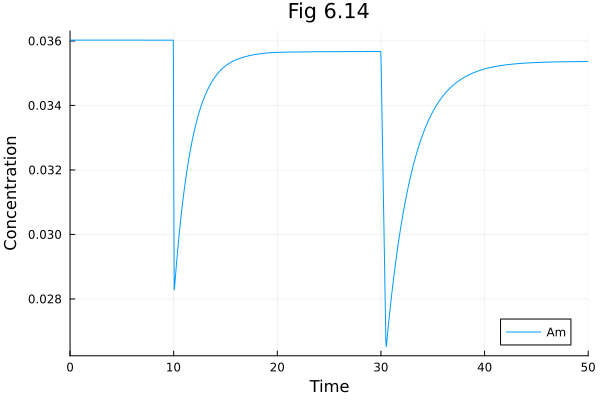
Fig 6.16¶
Model of apoptosis signalling pathway
using Catalyst
using OrdinaryDiffEq
using Plots
@time "Build system" rn616 = @reaction_network begin
@discrete_events begin
(t == 100.0) => [I => 200.0]
(t == 1200.0) => [I => 0.0]
end
(k1, k2), 0 <--> C8
k3 * (C3s + I), C8 --> C8s
k4, C8s --> 0
(k5, k6), C8s + BAR <--> C8sBAR
(k7, k8), 0 <--> C3
k9 * C8s, C3 --> C3s
k10, C3s --> 0
(k11, k12), C3s + IAP <--> C3sIAP
(k13, k14), 0 <--> BAR
(k15, k16 + k17 * C3s), 0 <--> IAP
k18, C8sBAR --> 0
k19, C3sIAP --> 0
endBuild system: 0.373275 seconds (1.36 M allocations: 67.656 MiB, 99.57% compilation time: 44% of which was recompilation)
Loading...
up616 = Dict(
:C8 => 1.3E5,
:C8s => 0.0,
:C3 => 0.21E5,
:C3s => 0.0,
:BAR => 0.4E5,
:IAP => 0.4E5,
:C8sBAR => 0.0,
:C3sIAP => 0.0,
:k1 => 507.0,
:k2 => 3.9e-3,
:k3 => 1e-5,
:k4 => 5.8e-3,
:k5 => 5e-4,
:k6 => 0.21,
:k7 => 81.9,
:k8 => 3.9e-3,
:k9 => 5.8e-6,
:k10 => 5.8e-3,
:k11 => 5e-4,
:k12 => 0.21,
:k13 => 40.0,
:k14 => 1e-3,
:k15 => 464.0,
:k16 => 1.16e-2,
:k17 => 3e-4,
:k18 => 1.16e-2,
:k19 => 1.73e-2,
:I => 0.0,
)
tend = 1800.0
@time "Build problem" prob616 = ODEProblem(rn616, up616, (0.0, tend); remove_conserved=true)
@time "Solve problem" sol616 = solve(prob616, FBDF(); tstops=[100.0, 1200.0])
@unpack C3s, C8s, I = rn616
plot(sol616, idxs=[C3s, C8s, I*100], title="Fig 6.16", xlabel="Time", ylabel="Concentration", legend=:right)Build problem: 2.821164 seconds (5.25 M allocations: 272.095 MiB, 0.97% gc time, 98.53% compilation time: 10% of which was recompilation)
Solve problem: 2.799010 seconds (5.83 M allocations: 298.644 MiB, 0.93% gc time, 99.66% compilation time)
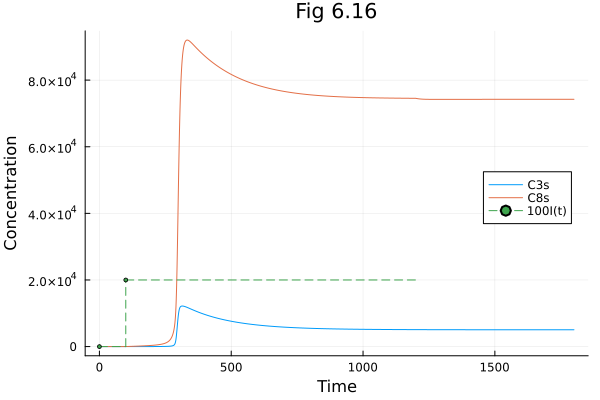
Fig 6.18¶
Model of calcium-induced calcium release in hepatocytes
using Catalyst
using OrdinaryDiffEq
using Plots
@time "Build system" rn618 = @reaction_network begin
@discretes I(t)
(k1 * I, km1), R <--> RI
(k2 * C, km2), RI <--> RIC
(k3 * C, km3), RIC <--> RICC
vr * (γ0 + γ1 * RIC) * (Cer - C), 0 --> C
hill(C, p1, p2, 4), C => 0
endBuild system: 0.344856 seconds (620.32 k allocations: 31.000 MiB, 99.59% compilation time: 3% of which was recompilation)
Loading...
up618 = Dict(
:C => 0.0,
:R => 1.0,
:RI => 0.0,
:RIC => 0.0,
:RICC => 0.0,
:k1 => 12.0,
:km1 => 8.0,
:k2 => 15.0,
:km2 => 1.65,
:k3 => 1.8,
:km3 => 0.21,
:vr => 0.185,
:γ0 => 0.1,
:γ1 => 20.5,
:p1 => 8.5,
:p2 => 0.065,
:Cer => 8.37,
:I => 0.0
)Dict{Symbol, Float64} with 18 entries:
:RI => 0.0
:RIC => 0.0
:R => 1.0
:p2 => 0.065
:k1 => 12.0
:km3 => 0.21
:k3 => 1.8
:vr => 0.185
:γ0 => 0.1
:γ1 => 20.5
:p1 => 8.5
:I => 0.0
:Cer => 8.37
:RICC => 0.0
:km1 => 8.0
:k2 => 15.0
:km2 => 1.65
:C => 0.0Fig 6.18 (A)¶
tspan = (0.0, 25.0)
up618a = merge(up618, Dict(:I => 2.0))
@time "Build problem" prob618a = ODEProblem(rn618, up618a, tspan; remove_conserved=true)
@time "Solve problem" sol618a = solve(prob618a, KenCarp47())
plot(sol618a, idxs=[:C, :RIC, :RICC], title="Fig 6.18 (A)", xlabel="Time", ylabel="Abundance", legend=:topright)Build problem: 1.244084 seconds (2.07 M allocations: 110.090 MiB, 97.69% compilation time)
Solve problem: 3.293846 seconds (7.59 M allocations: 399.372 MiB, 1.42% gc time, 99.71% compilation time)
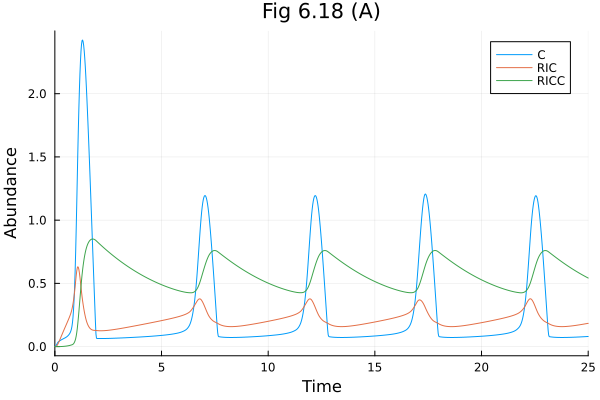
Fig 6.18 (B)¶
@unpack I = rn618
events = [
ModelingToolkitBase.SymbolicDiscreteCallback([20.0] => [I ~ 0.7]; discrete_parameters=[I]),
ModelingToolkitBase.SymbolicDiscreteCallback([60.0] => [I ~ 1.2]; discrete_parameters=[I]),
ModelingToolkitBase.SymbolicDiscreteCallback([90.0] => [I ~ 4.0]; discrete_parameters=[I])
]
osys618 = ode_model(rn618; remove_conserved=true, discrete_events=events)
tend = 120.0
prob618b = ODEProblem(osys618 |> complete, up618, tend)
@time sol618b = solve(prob618b, KenCarp47())
plot(sol618b, idxs=[:C], title="Fig 6.18 (B)", label="Calcium", xlabel="Time", ylabel="Concentration", legend=:topright) 7.693270 seconds (12.31 M allocations: 638.015 MiB, 0.63% gc time, 99.76% compilation time)
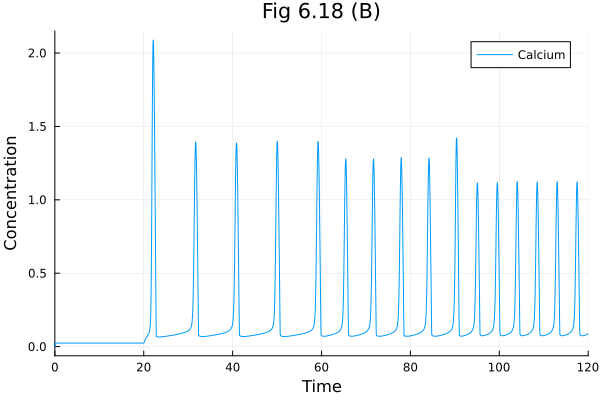
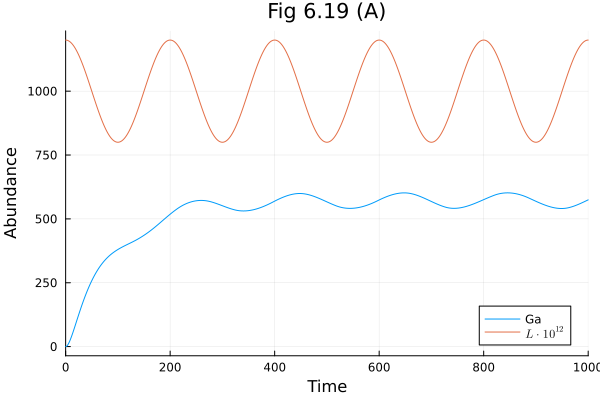
Fig 6.19¶
Sine wave response of g-protein signalling pathway
using OrdinaryDiffEq
using Catalyst
using Plots
using LaTeXStrings
@time "Build system" rn605 = @reaction_network begin
@parameters lt l_AMP l_per
@equations begin
L ~ lt + (lt / l_AMP) * cospi(2t / l_per)
end
kRL * L, R --> RL
kRLm, RL --> R
kGa, RL + G --> RL + Ga + Gbg
kGd0, Ga --> Gd
kG1, Gd + Gbg --> G
endBuild system: 0.445996 seconds (767.00 k allocations: 38.094 MiB, 99.48% compilation time: 2% of which was recompilation)
Loading...
up619 = Dict(
:kRL => 2e6,
:kRLm => 0.01,
:kGa => 1e-5,
:kGd0 => 0.11,
:kG1 => 1.0,
:R => 4e3,
:G => 1e4,
:lt => 1e-9,
:l_per => 200.0,
:l_AMP => 5.0,
:RL => 0.0,
:Ga => 0.0,
:Gd => 0.0,
:Gbg => 0.0
)
tend = 1000.0
@time "Build problem" prob619 = ODEProblem(rn605, up619, (0.0, tend); mtkcompile=true)
@time "Solve problem" sol619 = solve(prob619, KenCarp47())
@unpack Ga, L = rn605
plot(sol619, idxs=[Ga, L*1E12], title="Fig 6.19 (A)", xlabel="Time", ylabel="Abundance", label=["Ga" L"L \cdot 10^{12}"])Build problem: 1.495594 seconds (1.82 M allocations: 99.778 MiB, 98.24% compilation time: 49% of which was recompilation)
Solve problem: 0.771685 seconds (1.60 M allocations: 81.268 MiB, 3.02% gc time, 99.14% compilation time)
This notebook was generated using Literate.jl.