Fig 5.10, 5.11¶
Methionine model
using OrdinaryDiffEq
using ModelingToolkit
using ModelingToolkit: t_nounits as t, D_nounits as D
using Plots
Plots.gr(linewidth=1.5)Plots.GRBackend()Convenience functions to plot gradients of the ODEs.
function meshgrid(xrange, yrange)
xx = [x for y in yrange, x in xrange]
yy = [y for y in yrange, x in xrange]
return xx, yy
end
"Get the gradients of the vector field of an ODE system at a grid of points in the state space."
function get_gradient(prob, xsym, ysym, xx, yy; t = nothing)
# The order of state variables (unknowns) in the ODE system is not guaranteed. So we may need to swap the order of x and y when calling ∂F.
swap_or_not(x, y; xidx=1) = xidx == 1 ? [x, y] : [y, x]
∂F = prob.f
ps = prob.p
sys = prob.f.sys
xidx = ModelingToolkit.variable_index(sys, xsym)
yidx = ModelingToolkit.variable_index(sys, ysym)
dx = map((x, y) -> ∂F(swap_or_not(x, y; xidx), ps, t)[xidx], xx, yy)
dy = map((x, y) -> ∂F(swap_or_not(x, y; xidx), ps, t)[yidx], xx, yy)
return (dx, dy)
end
function normalize_gradient(dx, dy, xscale, yscale; transform=cbrt)
maxdx = maximum(abs, dx)
maxdy = maximum(abs, dy)
dx_norm = @. transform(dx / maxdx) * xscale
dy_norm = @. transform(dy / maxdy) * yscale
return (dx_norm, dy_norm)
endnormalize_gradient (generic function with 1 method)function model510(; name=:model510, simplfy=true)
hil(x, k=one(x)) = x / (x + k)
hil(x, k, n) = hil(x^n, k^n)
@parameters begin
K_AHC= 0.1
Adenosine = 1.0
v_MATI_max = 561.0
Met = 48.5
K_MATI_m = 41.0
K_MATI_i = 50.0
v_MATIII_max = 22870.0
K_MATIII_m2 = 21.1
v_GNMT_max = 10600.0
K_GNMT_m = 4500.0
K_GNMT_i = 20.0
v_MET_max = 4544.0
A_over_K_MET_m2 = 0.1
alpha_d = 1333.0
end
@variables AdoMet(t)=10 AdoHcy(t)=10
@variables Hcy(t) v_MATI(t) v_MATIII(t) K_MATIII_m1(t) v_GNMT(t) K_MET_m1(t) v_MET(t) v_D(t)
eqs = [
Hcy ~ AdoHcy * K_AHC / Adenosine,
v_MATI ~ v_MATI_max * hil(Met * hil(K_MATI_i, AdoMet), K_MATI_m),
v_MATIII ~ v_MATIII_max * hil(Met, K_MATIII_m1 * hil(K_MATIII_m2, Met)),
K_MATIII_m1 ~ 20000 / (1 + 5.7 * hil(AdoMet, 600)^2),
v_GNMT ~ v_GNMT_max * hil(AdoMet, K_GNMT_m, 2.3) * hil(K_GNMT_i, AdoHcy),
K_MET_m1 ~ 10 + 2.5 * AdoHcy,
v_MET ~ v_MET_max * hil(AdoMet, K_MET_m1) * hil(A_over_K_MET_m2),
v_D ~ alpha_d * Hcy,
D(AdoMet) ~ (v_MATI + v_MATIII) - (v_GNMT + v_MET),
D(AdoHcy) ~ (v_GNMT + v_MET - v_D) * hil(Adenosine, K_AHC)
]
sys = ODESystem(eqs, t; name)
return mtkcompile(sys; simplify)
endmodel510 (generic function with 1 method)Figure 5.10¶
@time "Build system" sys = model510()Build system: 45.953556 seconds (81.17 M allocations: 3.977 GiB, 1.96% gc time, 99.94% compilation time: 7% of which was recompilation)
Model model510:
Equations (2):
2 standard: see equations(model510)
Unknowns (2): see unknowns(model510)
AdoHcy(t)
AdoMet(t)
Parameters (14): see parameters(model510)
K_AHC
Adenosine
v_MATI_max
Met
⋮
Observed (8): see observed(model510)tend = 5.0
@time "Build problem" prob510 = ODEProblem(sys, [], (0.0, tend))
@time "Solve problem" sol510 = solve(prob510, KenCarp47())
plot(sol510, title="Fig. 5.10", xlabel="Time (s)", ylabel="Concentration (μM)")Build problem: 21.402789 seconds (55.34 M allocations: 2.741 GiB, 2.18% gc time, 99.85% compilation time: 55% of which was recompilation)
Solve problem: 2.748907 seconds (6.56 M allocations: 342.907 MiB, 1.28% gc time, 99.75% compilation time: <1% of which was recompilation)
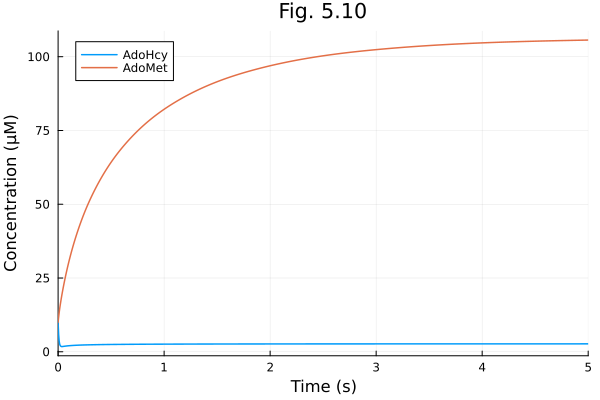
Figure 5.11 A¶
xrange = range(0, 1200, 101)
yrange = range(0, 6, 101)
xx, yy = meshgrid(xrange, yrange)
@unpack AdoMet, AdoHcy = prob510.f.sys
dx, dy = get_gradient(prob510, AdoMet, AdoHcy, xx, yy)
xx_sp = xx[1:5:end, 1:5:end]
yy_sp = yy[1:5:end, 1:5:end]
dx_sp = dx[1:5:end, 1:5:end]
dy_sp = dy[1:5:end, 1:5:end]
dx_sp_norm, dy_sp_norm = normalize_gradient(dx_sp, dy_sp, step(xrange) * 4 , step(yrange) * 4)([48.0 19.295821218268497 … -35.80742622518953 -37.93235774864831; 48.0 20.03422734530685 … -35.408918858013124 -37.53749940700541; … ; 48.0 27.524577217661925 … -28.41066989081311 -30.73950265642526; 48.0 27.78277653822824 … -28.015598637203375 -30.365908776833507], [0.0 0.17636544672548762 … 0.23526699058702708 0.24; -0.08520927651943645 0.16879128528066736 … 0.2308383769787323 0.23567946300920545; … ; -0.22737257395086835 -0.1930733128334119 … -0.06643736200699227 0.06946053098627193; -0.23129356128166506 -0.1987940355791439 … -0.09919332784830667 -0.07078013719696569])quiver(xx_sp, yy_sp, quiver=(dx_sp_norm, dy_sp_norm), color=:gray)
contour!(xrange, yrange, dx, levels=[0], line=(:black, :solid), colorbar=false)
plot!([], [], line=(:black, :solid), label="AdoMet nullcline")
contour!(xrange, yrange, dy, levels=[0], line=(:black, :dash), colorbar=false)
plot!([], [], line=(:black, :dash), label="AdoHcy nullcline")
plot!(title="Fig. 5.11 (A)", xlabel="AdoMet (μM)", ylabel="AdoHcy (μM)", legend=:bottomright, size=(800, 600), xlims=(0, 1200), ylims=(0, 6))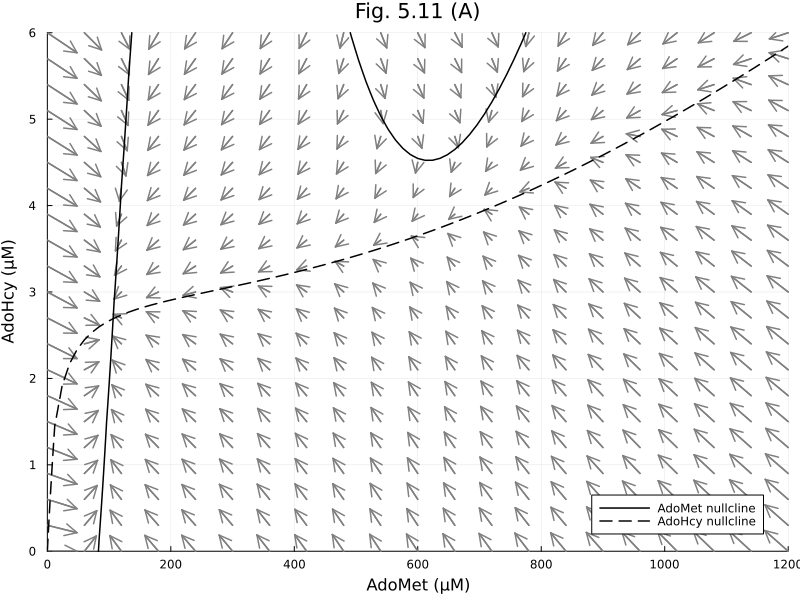
Figure 5.11 B¶
Increased methionine level
prob511b = remake(prob510, p=[sys.Met => 51.0])
xrange = range(0, 1200, 101)
yrange = range(0, 6, 101)
xx, yy = meshgrid(xrange, yrange)
@unpack AdoMet, AdoHcy = prob511b.f.sys
dx, dy = get_gradient(prob511b, AdoMet, AdoHcy, xx, yy)
xx_sp = xx[1:5:end, 1:5:end]
yy_sp = yy[1:5:end, 1:5:end]
dx_sp = dx[1:5:end, 1:5:end]
dy_sp = dy[1:5:end, 1:5:end]
dx_sp_norm, dy_sp_norm = normalize_gradient(dx_sp, dy_sp, step(xrange) * 4 , step(yrange) * 4)([48.0 22.84321705890981 … -31.74002305152799 -34.196483214992256; 48.0 23.354186304817638 … -31.253785820679994 -33.73117080825976; … ; 48.0 29.227915239078932 … -21.571870212890552 -24.99392261316023; 48.0 29.4469285093872 … -20.90640044972195 -24.44918451737791], [0.0 0.17636544672548762 … 0.23526699058702708 0.24; -0.08520927651943645 0.16879128528066736 … 0.2308383769787323 0.23567946300920545; … ; -0.22737257395086835 -0.1930733128334119 … -0.06643736200699227 0.06946053098627193; -0.23129356128166506 -0.1987940355791439 … -0.09919332784830667 -0.07078013719696569])quiver(xx_sp, yy_sp, quiver=(dx_sp_norm, dy_sp_norm), color=:gray)
contour!(xrange, yrange, dx, levels=[0], line=(:black, :solid), colorbar=false)
plot!([], [], line=(:black, :solid), label="AdoMet nullcline")
contour!(xrange, yrange, dy, levels=[0], line=(:black, :dash), colorbar=false)
plot!([], [], line=(:black, :dash), label="AdoHcy nullcline")
plot!(title="Fig. 5.11 (B)", xlabel="AdoMet (μM)", ylabel="AdoHcy (μM)", legend=:bottomright, size=(800, 600), xlims=(0, 1200), ylims=(0, 6))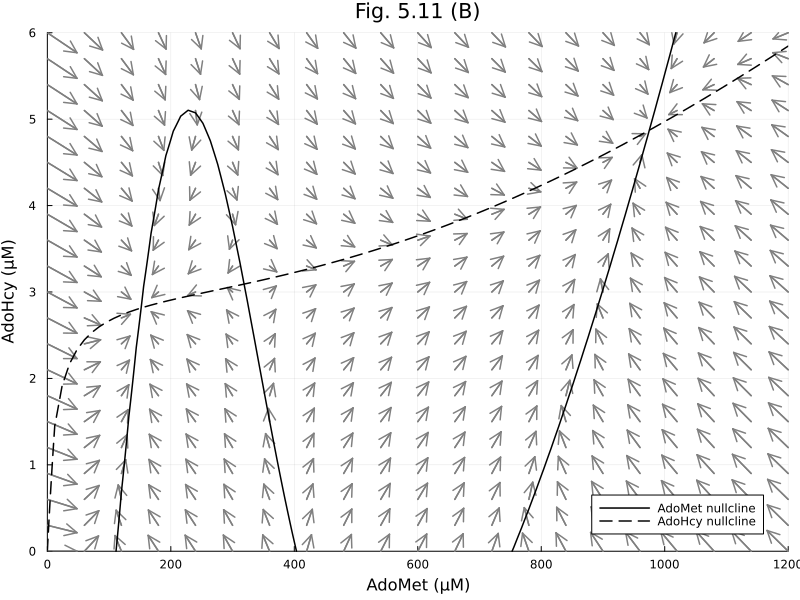
This notebook was generated using Literate.jl.