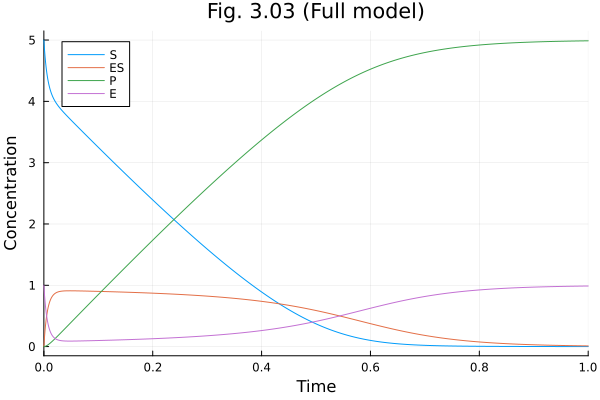
Figure 3.03¶
Michaelis-Menten kinetics
using OrdinaryDiffEq
using Catalyst
using PlotsEnzyme kinetics full model
@time "Build system" rn303 = @reaction_network begin
k1, S + E --> ES
km1, ES --> S + E
k2, ES --> P + E
endBuild system: 0.553776 seconds (4.74 M allocations: 237.683 MiB, 4.27% gc time, 99.64% compilation time)
Loading...
up303 = Dict(:S => 5.0, :ES => 0.0, :P => 0.0, :E=>1.0, :k1 => 30.0, :km1 => 1.0, :k2 => 10.0)
tend = 1.0
@time "Build problem" prob303 = ODEProblem(rn303, up303, (0.0, tend); remove_conserved=true)Build problem: 10.765757 seconds (40.39 M allocations: 1.973 GiB, 7.55% gc time, 99.75% compilation time: 70% of which was recompilation)
ODEProblem with uType Vector{Float64} and tType Float64. In-place: true
Initialization status: FULLY_DETERMINED
Non-trivial mass matrix: false
timespan: (0.0, 1.0)
u0: 2-element Vector{Float64}:
5.0
1.0@time "Solve problem" sol303 = solve(prob303, Tsit5())
plot(sol303, idxs=[:S, :ES, :P, :E], title="Fig. 3.03 (Full model)", xlabel="Time", ylabel="Concentration")Solve problem: 1.291483 seconds (6.26 M allocations: 330.051 MiB, 2.04% gc time, 99.86% compilation time: <1% of which was recompilation)
QSSA of the ES complex¶
@time "Build system" rn303mm = @reaction_network begin
mm(S, k2 * ET, (km1 + k2) / k1), S => P
endBuild system: 0.039047 seconds (329.12 k allocations: 16.378 MiB, 97.97% compilation time)
Loading...
up303mm = Dict(:k1 => 30.0, :km1 => 1.0, :k2 => 10.0, :ET => 1.0, :S => 5.0, :P => 0.0)
tend = 1.0
@time "Build problem" prob303mm = ODEProblem(rn303mm, up303mm, (0.0, tend); remove_conserved=true)
@time "Solve problem" sol303mm = solve(prob303mm, Tsit5())
plot(sol303mm, idxs=[:S, :P], labels=["S (QSSA)" "P (QSSA)"], title="Fig. 3.03 (QSSA vs Full)", xlabel="Time", ylabel="Concentration")
plot!(sol303, idxs=[:S, :P], labels=["S (full)" "P (full)"], linestyle=:dash, legend=:right)Build problem: 0.582577 seconds (2.39 M allocations: 127.014 MiB, 97.56% compilation time)
Solve problem: 0.406917 seconds (2.15 M allocations: 109.323 MiB, 5.87% gc time, 99.50% compilation time)
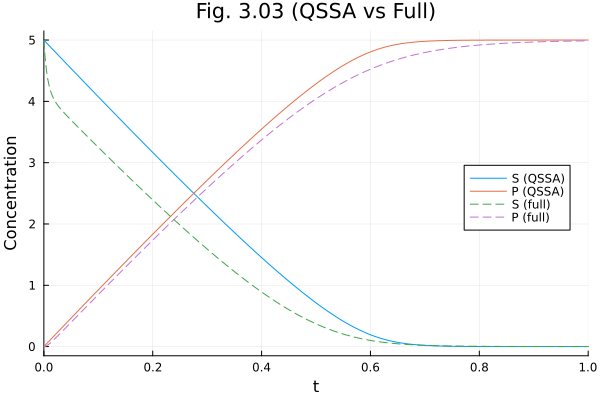
Fig 3.13¶
Generalized mass action (GMA) vs. Michaelis-Menten rate laws
using Plots
plot([t -> 2t / (1+t), t -> t^0.4], 0, 4, labels=["MM" "GMA"], title="Fig. 3.13", xlabel="Substrate concentration (S / Km)", ylabel="Reaction rate (AU)")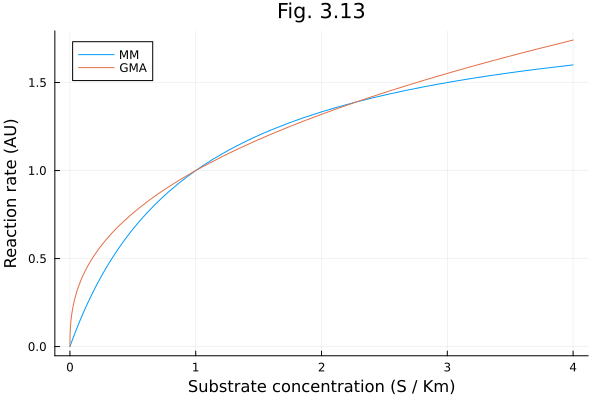
This notebook was generated using Literate.jl.