Fig 2.04¶
Exponential decay
using Plots
Plots.default(linewidth=1.5)pl = plot(xlabel="Time", ylabel="Concentration", title="Fig 2.4")
for k in 1:3
plot!(pl, t -> 3 * exp(-k * t), 0, 5, label="exp(-$(k)t)")
end
pl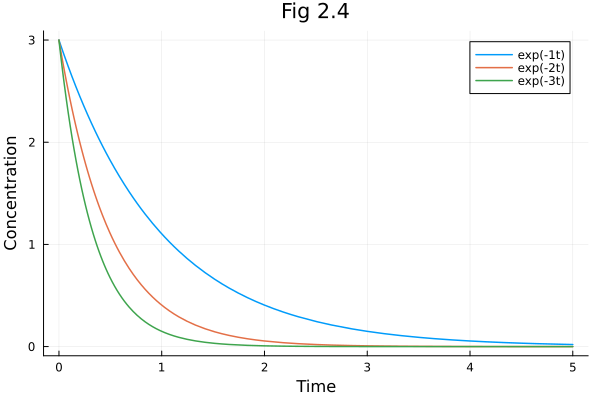
Fig 2.09¶
Metabolic network simulation
using OrdinaryDiffEq
using Catalyst
using Plots@time "Build system" rn209 = @reaction_network model209 begin
k1, 0 --> A
k2, A --> B
k3, A + B --> C + D
k4, C --> 0
k5, D --> 0
endBuild system: 0.924045 seconds (5.29 M allocations: 264.458 MiB, 32.86% gc time, 99.79% compilation time)
Loading...
up209 = (:k1 => 3.0, :k2 => 2.0, :k3 => 2.5, :k4 => 3.0, :k5 => 4.0, :A => 0.0, :B => 0.0, :C => 0.0, :D => 0.0)
tend = 10.0
@time "Build problem" prob209 = ODEProblem(rn209, up209, tend)
@time "Solve problem" sol = solve(prob209, Tsit5())
plot(sol, title="Fig 2.9", xlabel="Time", ylabel="Concentration")Build problem: 10.741642 seconds (42.03 M allocations: 2.048 GiB, 4.51% gc time, 99.53% compilation time: 64% of which was recompilation)
Solve problem: 1.451551 seconds (6.94 M allocations: 363.527 MiB, 1.68% gc time, 99.82% compilation time)
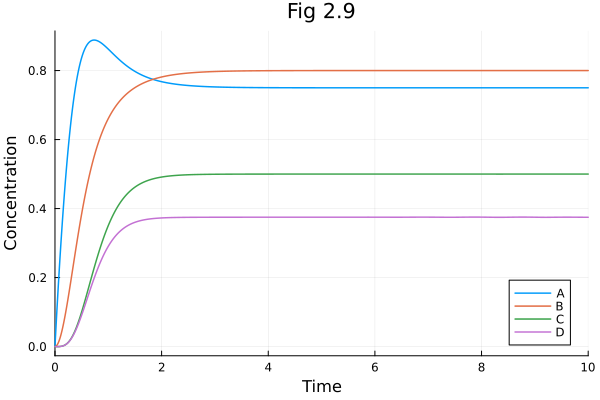
Figure 2.11¶
Intro to model reduction of ODE metabolic networks.
using OrdinaryDiffEq
using Catalyst
using Plots@time "Build system" rn211 = @reaction_network model211 begin
k0, 0 --> A
(k1, km1), A <--> B
k2, B --> 0
endBuild system: 0.000370 seconds (1.36 k allocations: 51.516 KiB)
Loading...
up211 = (:k0 => 0.0, :k1 => 9.0, :km1 => 12.0, :k2 => 2.0, :A => 0.0, :B => 10.0)
tend = 3.0
@time "Build problem" prob211 = ODEProblem(rn211, up211, tend)
@time "Solve problem" sol211 = solve(prob211, Tsit5())
plot(sol211, title="Fig 2.11", xlabel="Time", ylabel="Concentration")Build problem: 0.637793 seconds (3.49 M allocations: 175.044 MiB, 2.87% gc time, 98.30% compilation time)
Solve problem: 0.348996 seconds (1.60 M allocations: 81.761 MiB, 99.41% compilation time)
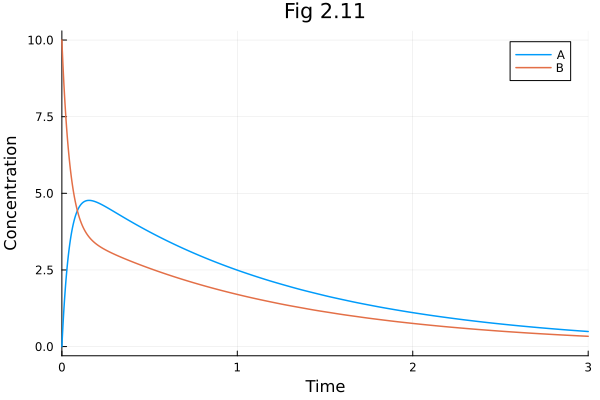
Figure 2.12¶
Rapid equilibrium assumption
@time "Build system" rn212 = @reaction_network model212 begin
@parameters k0 k1 km1 k2
@equations begin
A ~ C * km1 / (km1 + k1)
B ~ C * k1 / (km1 + k1)
end
k0, 0 --> C
k2 * k1 / (km1 + k1), C --> 0
endBuild system: 0.177102 seconds (474.93 k allocations: 23.551 MiB, 99.34% compilation time)
Loading...
tend = 3.0
up212 = (:k0 => 0.0, :k1 => 9.0, :km1 => 12.0, :k2 => 2.0, :C => 10.0)
@time "Build problem" prob212 = ODEProblem(rn212, up212, tend; mtkcompile=true)
@time "Solve problem" sol212 = solve(prob212, Tsit5())
plot(sol211, linestyle=:dash, label=["A (Full model)" "B (Full model)"])
plot!(sol212, idxs=[:A, :B], linestyle=:solid, label=["A (Reduced model)" "B (Reduced model)"])
plot!(title="Fig 2.12", xlabel="Time", ylabel="Concentration")Build problem: 0.981062 seconds (3.94 M allocations: 195.815 MiB, 2.14% gc time, 98.76% compilation time: 35% of which was recompilation)
Solve problem: 0.351015 seconds (1.63 M allocations: 83.084 MiB, 99.45% compilation time)
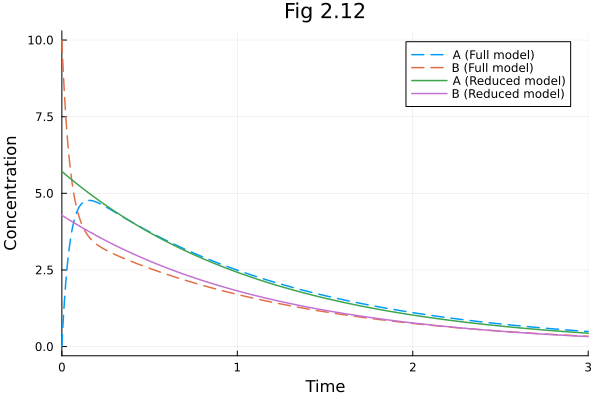
Figure 2.13¶
When another set of parameters breaks rapid equilibrium assumption.
pmap213 = Dict(:k0=>9.0, :k1=>21.0, :km1=>12.0, :k2=>2.0)
u0map213 = Dict(:A=>8.0, :B=>4.0)
tend = 3.0
prob213full = remake(prob211; p=pmap213, u0=u0map213, tspan=(0.0, tend))
prob213re = ODEProblem(rn212, (:C=>12.0), tend, pmap213; mtkcompile=true)
@time sol213full = solve(prob213full, Tsit5())
@time sol213re = solve(prob213re, Tsit5())
plot(sol213full, linestyle=:dash, label=["A (Full model)" "B (Full model)"])
plot!(sol213re, idxs=[:A, :B], linestyle=:solid, label=["A (Reduced model)" "B (Reduced model)"])
plot!(title="Fig 2.13", xlabel="Time", ylabel="Concentration") 0.000216 seconds (950 allocations: 47.445 KiB)
0.000200 seconds (464 allocations: 25.680 KiB)
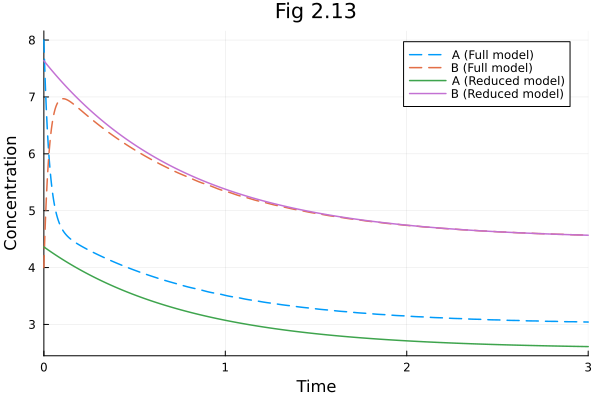
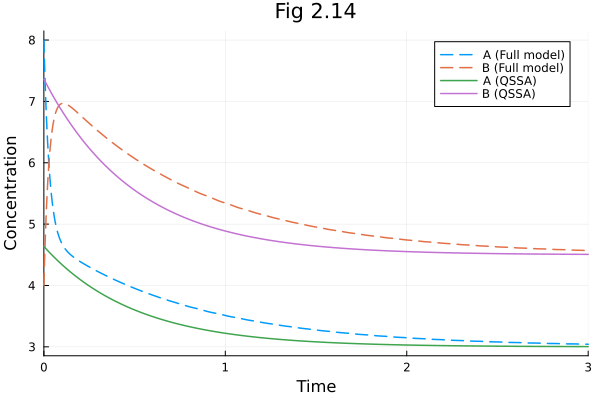
Figure 2.14¶
Quasi-steady state assumption on species A (D(A) ~ 0). You can model this in Catalyst.jl DSL like:
@time "Build system" rn214 = @reaction_network model214 begin
@parameters k0=9 k1=21 km1=12 k2=2
@variables A(t)
@equations begin
A ~ (k0 + km1 * B) / k1
end
(k1 * A, (k2 + km1)), 0 <--> B
endBuild system: 0.022769 seconds (46.52 k allocations: 2.294 MiB, 96.44% compilation time)
Loading...
Or use ModelingToolkit.jl equations to set dA ~ 0 directly. The QSS concentrations of A
using ModelingToolkit
function model214(u0; name=:model214)
@independent_variables t
@parameters k0=9 k1=21 km1=12 k2=2
D = Differential(t)
@variables A(t) B(t) = (k1 * u0 - k0) / (km1 + k1)
v0 = k0
v1 = k1 * A - km1 * B
v2 = k2 * B
dA = v0 - v1
dB = v1 - v2
eqs = [
0 ~ dA, ## Quasi-steady state assumption on species A (`D(A) ~ 0`)
D(B) ~ dB,
]
return ODESystem(eqs, t; name)
end
tend = 3.0
@time "Build system" @mtkcompile sys214 = model214(12.0)
@time "Build problem" prob214 = ODEProblem(sys214, [], (0.0, tend))Build system: 0.455626 seconds (3.55 M allocations: 174.693 MiB, 99.57% compilation time: <1% of which was recompilation)
Build problem: 0.442491 seconds (1.42 M allocations: 78.201 MiB, 4.52% gc time, 97.81% compilation time: 10% of which was recompilation)
ODEProblem with uType Vector{Float64} and tType Float64. In-place: true
Initialization status: FULLY_DETERMINED
Non-trivial mass matrix: false
timespan: (0.0, 3.0)
u0: 1-element Vector{Float64}:
7.363636363636363equations(sys214)Loading...
observed(sys214)Loading...
@time "Solve problem" sol214 = solve(prob214, Tsit5())
@unpack A, B = sys214
plot(sol213full, linestyle=:dash, label=["A (Full model)" "B (Full model)"])
plot!(sol214, idxs=[A, B], linestyle=:solid, label=["A (QSSA)" "B (QSSA)"])
plot!(title="Fig 2.14", xlabel="Time", ylabel="Concentration")Solve problem: 0.259810 seconds (1.27 M allocations: 65.422 MiB, 99.32% compilation time)
Problem 2.4.6¶
using OrdinaryDiffEq
using Plots
sol246 = ODEProblem((u, p, t) -> p * (1.0 - u), 0.0, 10.0, 1.0) |> solve
plot(sol246, title="Problem 2.4.6", xlabel="Time", ylabel="Concentration")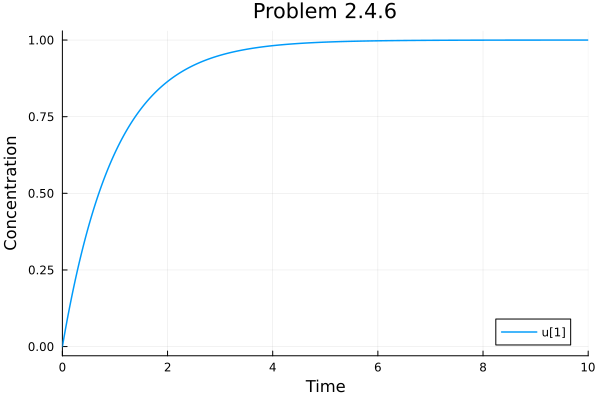
This notebook was generated using Literate.jl.