Standard procedures
Define a model function representing the right-hand-side (RHS) of the system.
Out-of-place form:
f(u, p, t)whereuis the state variable(s),pis the parameter(s), andtis the independent variable (usually time). The output is the right hand side (RHS) of the differential equation system.In-place form:
f!(du, u, p, t), where the output is saved todu. The rest is the same as the out of place form. The in-place form has potential performance benefits since it allocates less than the out-of-place (f(u, p, t)) counterpart.Using ModelingToolkit.jl : define equations and build an ODE system.
Initial conditions (
u0) for the state variable(s).(Optional) define parameter(s)
p.Define a problem (e.g.
ODEProblem) using the modeling function (f), initial conditions (u0), simulation time span (tspan == (tstart, tend)), and parameter(s)p.Solve the problem by calling
solve(prob).
Solve ODEs using OrdinaryDiffEq.jl¶
Documentation: https://
Single variable: Exponential decay model¶
The concentration of a decaying nuclear isotope could be described as an exponential decay:
State variable
: The concentration of a decaying nuclear isotope.
Parameter
: The rate constant of decay. The half-life
using OrdinaryDiffEq
using PlotsThe model function is the 3-argument out-of-place form, f(u, p, t).
decay(u, p, t) = p * u
p = -1.0 ## Rate of exponential decay
u0 = 1.0 ## Initial condition
tspan = (0.0, 2.0) ## Start time and end time
prob = ODEProblem(decay, u0, tspan, p)
sol = solve(prob)retcode: Success
Interpolation: 3rd order Hermite
t: 8-element Vector{Float64}:
0.0
0.10001999200479662
0.34208427066999536
0.6553980136343391
1.0312652525315806
1.4709405856363595
1.9659576669700232
2.0
u: 8-element Vector{Float64}:
1.0
0.9048193287657775
0.7102883621328674
0.5192354400036404
0.3565557657699655
0.22970979078638265
0.1400224727245278
0.13533600284008826Solution at time t=1.0 (with interpolation)
sol(1.0)0.3678796381978344Time points
sol.t8-element Vector{Float64}:
0.0
0.10001999200479662
0.34208427066999536
0.6553980136343391
1.0312652525315806
1.4709405856363595
1.9659576669700232
2.0Solutions at sol.t
sol.u8-element Vector{Float64}:
1.0
0.9048193287657775
0.7102883621328674
0.5192354400036404
0.3565557657699655
0.22970979078638265
0.1400224727245278
0.13533600284008826Visualize the solution
plot(sol, title="Exponential Decay", xlabel="Time", ylabel="Concentration")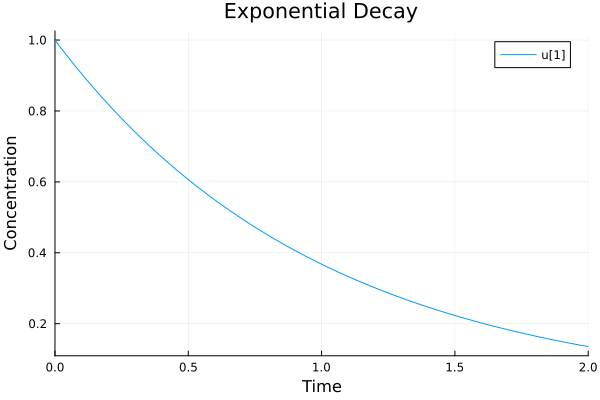
Three variables: The SIR model¶
The SIR model describes the spreading of an contagious disease can be described by the SIR model:
State variables
: the fraction of susceptible people
: the fraction of infectious people
: the fraction of recovered (or removed) people
Parameters
: the rate of infection when susceptible and infectious people meet
: the rate of recovery of infectious people
using OrdinaryDiffEq
using PlotsSIR model (in-place form can save array allocations and thus faster)
function sir!(du, u, p, t)
s, i, r = u
β, γ = p
v1 = β * s * i
v2 = γ * i
du[1] = -v1
du[2] = v1 - v2
du[3] = v2
return nothing
endsir! (generic function with 1 method)p = (β=1.0, γ=0.3)
u0 = [0.99, 0.01, 0.00]
tspan = (0.0, 20.0)
prob = ODEProblem(sir!, u0, tspan, p)
sol = solve(prob)retcode: Success
Interpolation: 3rd order Hermite
t: 17-element Vector{Float64}:
0.0
0.08921318693905476
0.3702862715172094
0.7984257132319627
1.3237271485666187
1.991841832691831
2.7923706947355837
3.754781614278828
4.901904318934307
6.260476636498209
7.7648912410433075
9.39040980993922
11.483861023017885
13.372369854616487
15.961357172044833
18.681426667664056
20.0
u: 17-element Vector{Vector{Float64}}:
[0.99, 0.01, 0.0]
[0.9890894703413342, 0.010634484617786016, 0.00027604504087978485]
[0.9858331594901347, 0.012901496825852227, 0.0012653436840130792]
[0.9795270529591532, 0.017282420996456258, 0.003190526044390598]
[0.9689082167415561, 0.024631267034445452, 0.006460516223998509]
[0.9490552312363142, 0.03827338797605376, 0.012671380787632129]
[0.911862947533394, 0.06347250098224955, 0.024664551484356523]
[0.8398871089274514, 0.11078176031568526, 0.04933113075686341]
[0.707584206802473, 0.19166147882272797, 0.10075431437479916]
[0.5081460287219878, 0.2917741934147055, 0.2000797778633069]
[0.31213222024414106, 0.3415879120018043, 0.34627986775405484]
[0.1821568309636563, 0.309998313415639, 0.5078448556207048]
[0.10427205468919257, 0.2206111401113331, 0.6751168051994745]
[0.07386737407725885, 0.14760143051851196, 0.7785311954042293]
[0.05545028910907742, 0.07997076922865369, 0.864578941662269]
[0.04733499069589219, 0.040605653213833574, 0.9120593560902743]
[0.04522885458929347, 0.02905741611081477, 0.9257137292998918]Visualize the solution
plot(sol, title="SIR model", xlabel="Time", ylabel="Fraction of population", labels=["S" "I" "R"])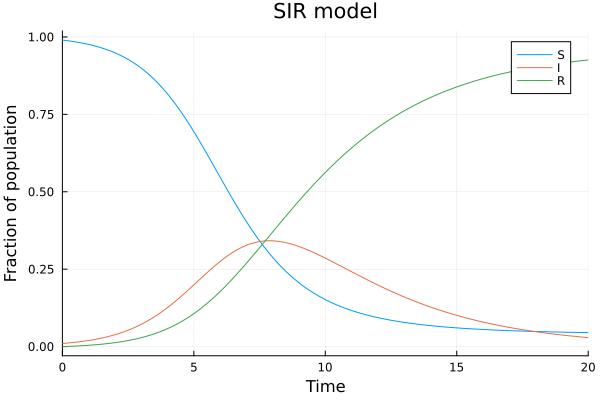
Saving simulation results¶
using DataFrames
using CSV
df = DataFrame(sol)
CSV.write("sir.csv", df)
rm("sir.csv")This notebook was generated using Literate.jl.